Competing Pathways and Multiple Folding Nuclei in a Large Multidomain Protein, Luciferase
- PMID: 28494954
- PMCID: PMC5425382
- DOI: 10.1016/j.bpj.2017.03.028
Competing Pathways and Multiple Folding Nuclei in a Large Multidomain Protein, Luciferase
Abstract
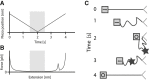
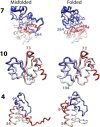
Proteins obtain their final functional configuration through incremental folding with many intermediate steps in the folding pathway. If known, these intermediate steps could be valuable new targets for designing therapeutics and the sequence of events could elucidate the mechanism of refolding. However, determining these intermediate steps is hardly an easy feat, and has been elusive for most proteins, especially large, multidomain proteins. Here, we effectively map part of the folding pathway for the model large multidomain protein, Luciferase, by combining single-molecule force-spectroscopy experiments and coarse-grained simulation. Single-molecule refolding experiments reveal the initial nucleation of folding while simulations corroborate these stable core structures of Luciferase, and indicate the relative propensities for each to propagate to the final folded native state. Both experimental refolding and Monte Carlo simulations of Markov state models generated from simulation reveal that Luciferase most often folds along a pathway originating from the nucleation of the N-terminal domain, and that this pathway is the least likely to form nonnative structures. We then engineer truncated variants of Luciferase whose sequences corresponded to the putative structure from simulation and we use atomic force spectroscopy to determine their unfolding and stability. These experimental results corroborate the structures predicted from the folding simulation and strongly suggest that they are intermediates along the folding pathway. Taken together, our results suggest that initial Luciferase refolding occurs along a vectorial pathway and also suggest a mechanism that chaperones may exploit to prevent misfolding.
Copyright © 2017 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures





Similar articles
-
Chaperones rescue luciferase folding by separating its domains.J Biol Chem. 2014 Oct 10;289(41):28607-18. doi: 10.1074/jbc.M114.582049. Epub 2014 Aug 26. J Biol Chem. 2014. PMID: 25160632 Free PMC article.
-
Misfolding of luciferase at the single-molecule level.Angew Chem Int Ed Engl. 2014 Sep 22;53(39):10390-3. doi: 10.1002/anie.201405566. Epub 2014 Aug 14. Angew Chem Int Ed Engl. 2014. PMID: 25124399
-
Characterization of the folding and unfolding reactions of single-chain monellin: evidence for multiple intermediates and competing pathways.Biochemistry. 2007 Oct 23;46(42):11727-43. doi: 10.1021/bi701142a. Epub 2007 Sep 29. Biochemistry. 2007. PMID: 17902706
-
Single-Molecule Force Spectroscopy of Protein Folding.J Mol Biol. 2021 Oct 1;433(20):167207. doi: 10.1016/j.jmb.2021.167207. Epub 2021 Aug 18. J Mol Biol. 2021. PMID: 34418422 Review.
-
Successes and challenges in simulating the folding of large proteins.J Biol Chem. 2020 Jan 3;295(1):15-33. doi: 10.1074/jbc.REV119.006794. Epub 2019 Nov 11. J Biol Chem. 2020. PMID: 31712314 Free PMC article. Review.
Cited by
-
Mechanical Stability of a Small, Highly-Luminescent Engineered Protein NanoLuc.Int J Mol Sci. 2020 Dec 23;22(1):55. doi: 10.3390/ijms22010055. Int J Mol Sci. 2020. PMID: 33374567 Free PMC article.
-
3β-Corner Stability by Comparative Molecular Dynamics Simulations.Int J Mol Sci. 2022 Oct 2;23(19):11674. doi: 10.3390/ijms231911674. Int J Mol Sci. 2022. PMID: 36232976 Free PMC article.
-
The Ribosome Cooperates with a Chaperone to Guide Multi-domain Protein Folding.Mol Cell. 2019 Apr 18;74(2):310-319.e7. doi: 10.1016/j.molcel.2019.01.043. Epub 2019 Mar 6. Mol Cell. 2019. PMID: 30852061 Free PMC article.
-
Mechanical Forces and Their Effect on the Ribosome and Protein Translation Machinery.Cells. 2020 Mar 7;9(3):650. doi: 10.3390/cells9030650. Cells. 2020. PMID: 32156009 Free PMC article. Review.
-
The conformational landscape of a serpin N-terminal subdomain facilitates folding and in-cell quality control.bioRxiv [Preprint]. 2023 Apr 26:2023.04.24.537978. doi: 10.1101/2023.04.24.537978. bioRxiv. 2023. PMID: 37163105 Free PMC article. Preprint.
References
-
- Gruebele M., Dave K., Sukenik S. Globular protein folding in vitro and in vivo. Annu. Rev. Biophys. 2016;45:233–251. - PubMed
-
- Dobson C.M. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Fersht A. W. H. Freeman; New York: 1999. Structure and Mechanism in Protein Science: A Guide to Enzyme Catalysis and Protein Folding.
-
- Dill K.A., MacCallum J.L. The protein-folding problem, 50 years on. Science. 2012;338:1042–1046. - PubMed
-
- Gruebele M. Protein folding: the free energy surface. Curr. Opin. Struct. Biol. 2002;12:161–168. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

