Elevated μs-ms timescale backbone dynamics in the transition state analog form of arginine kinase
- PMID: 28495594
- PMCID: PMC5677571
- DOI: 10.1016/j.jsb.2017.05.002
Elevated μs-ms timescale backbone dynamics in the transition state analog form of arginine kinase
Abstract
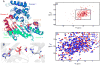
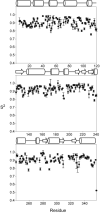
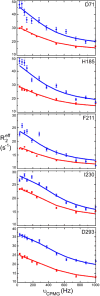
Arginine kinase catalyzes reversible phosphoryl transfer between arginine and ATP. Crystal structures of arginine kinase in an open, substrate-free form and closed, transition state analog (TSA) complex indicate that the enzyme undergoes substantial domain and loop rearrangements required for substrate binding, catalysis, and product release. Nuclear magnetic resonance (NMR) has shown that substrate-free arginine kinase is rigid on the ps-ns timescale (average S2=0.84±0.08) yet quite dynamic on the µs-ms timescale (35 residues with Rex, 12%), and that movements of the N-terminal domain and the loop comprising residues I182-G209 are rate-limiting on catalysis. Here, NMR of the TSA-bound enzyme shows similar rigidity on the ps-ns timescale (average S2=0.91±0.05) and substantially increased μs-ms timescale dynamics (77 residues; 22%). Many of the residues displaying μs-ms dynamics in NMR Carr-Purcell-Meiboom-Gill (CPMG) 15N backbone relaxation dispersion experiments of the TSA complex are also dynamic in substrate-free enzyme. However, the presence of additional dynamic residues in the TSA-bound form suggests that dynamics extend through much of the C-terminal domain, which indicates that in the closed form, a larger fraction of the protein takes part in conformational transitions to the excited state(s). Conformational exchange rate constants (kex) of the TSA complex are all approximately 2500s-1, higher than any observed in the substrate-free enzyme (800-1900s-1). Elevated μs-ms timescale protein dynamics in the TSA-bound enzyme is more consistent with recently postulated catalytic networks involving multiple interconnected states at each step of the reaction, rather than a classical single stabilized transition state.
Keywords: Arginine kinase; Dynamics; NMR; Relaxation dispersion; Transition state analog.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures

References
-
- Boehr DD, McElheny D, Dyson HJ, Wright PE. The dynamic energy landscape of dihydrofolate reductase catalysis. Science. 2006;313:1638–1642. - PubMed
-
- Borders CL, Jr, Snider MJ, Wolfenden R, Edmiston PL. Determination of the affinity of each component of a composite quaternary transition-state analogue complex of creatine kinase. Biochemistry. 2002;41:6995–7000. - PubMed
-
- Brath U, Akke M. Differential responses of the backbone and side-chain conformational dynamics in FKBP12 upon binding the transition-state analog FK506: implications for transition-state stabilization and target protein recognition. J Mol Biol. 2009;387:233–244. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

