Transcription factor-associated combinatorial epigenetic pattern reveals higher transcriptional activity of TCF7L2-regulated intragenic enhancers
- PMID: 28499350
- PMCID: PMC5429574
- DOI: 10.1186/s12864-017-3764-9
Transcription factor-associated combinatorial epigenetic pattern reveals higher transcriptional activity of TCF7L2-regulated intragenic enhancers
Abstract
Background: Recent studies have suggested that combinations of multiple epigenetic modifications are essential for controlling gene expression. Despite numerous computational approaches have been developed to decipher the combinatorial epigenetic patterns or "epigenetic code", none of them has explicitly addressed the relationship between a specific transcription factor (TF) and the patterns.
Methods: Here, we developed a novel computational method, T-cep, for annotating chromatin states associated with a specific TF. T-cep is composed of three key consecutive modules: (i) Data preprocessing, (ii) HMM training, and (iii) Potential TF-states calling.
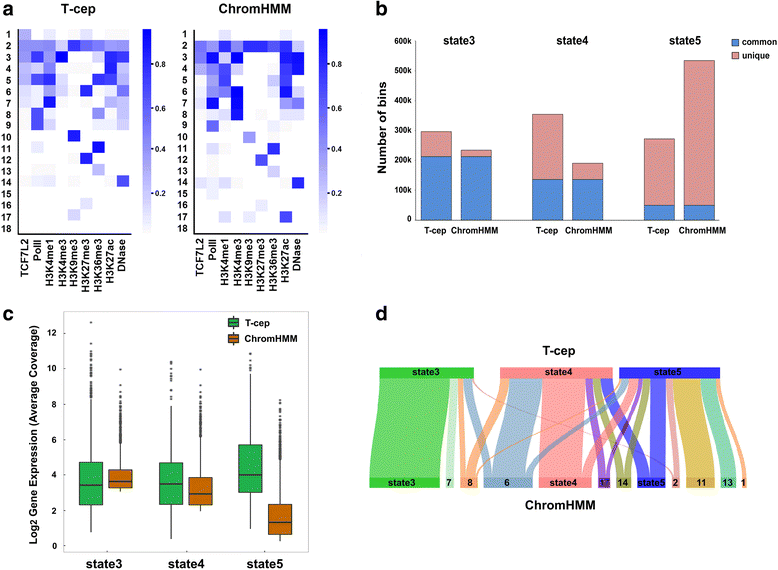
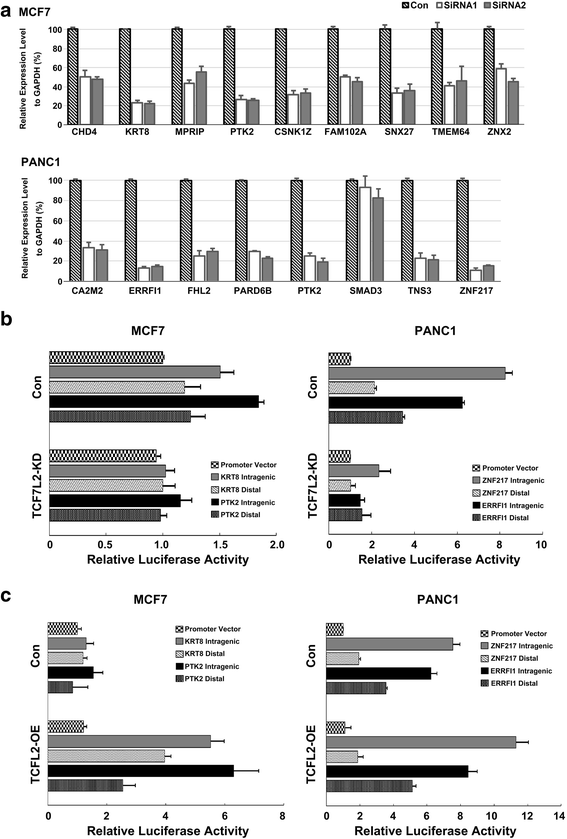
Results: We evaluated T-cep on a TCF7L2-omics data. Unexpectedly, our method has uncovered a novel set of TCF7L2-regulated intragenic enhancers missed by other software tools, where the associated genes exert the highest gene expression. We further used siRNA knockdown, Co-transfection, RT-qPCR and Luciferase Reporter Assay not only to validate the accuracy and efficiency of prediction by T-cep, but also to confirm the functionality of TCF7L2-regulated enhancers in both MCF7 and PANC1 cells respectively.
Conclusions: Our study for the first time at a genome-wide scale reveals the enhanced transcriptional activity of cell-type-specific TCF7L2 intragenic enhancers in regulating gene expression.
Keywords: Intragenic enhancer; MCF7; PANC1; T-cep; TCF7L2.
Figures



Similar articles
-
Cell type-specific binding patterns reveal that TCF7L2 can be tethered to the genome by association with GATA3.Genome Biol. 2012 Sep 26;13(9):R52. doi: 10.1186/gb-2012-13-9-r52. Genome Biol. 2012. PMID: 22951069 Free PMC article.
-
Enhancer decommissioning by Snail1-induced competitive displacement of TCF7L2 and down-regulation of transcriptional activators results in EPHB2 silencing.Biochim Biophys Acta. 2016 Nov;1859(11):1353-1367. doi: 10.1016/j.bbagrm.2016.08.002. Epub 2016 Aug 5. Biochim Biophys Acta. 2016. PMID: 27504909
-
Three-dimensional analysis reveals altered chromatin interaction by enhancer inhibitors harbors TCF7L2-regulated cancer gene signature.J Cell Biochem. 2019 Mar;120(3):3056-3070. doi: 10.1002/jcb.27449. Epub 2018 Dec 11. J Cell Biochem. 2019. PMID: 30548288 Free PMC article.
-
Towards genome-wide prediction and characterization of enhancers in plants.Biochim Biophys Acta Gene Regul Mech. 2017 Jan;1860(1):131-139. doi: 10.1016/j.bbagrm.2016.06.006. Epub 2016 Jun 16. Biochim Biophys Acta Gene Regul Mech. 2017. PMID: 27321818 Review.
-
Transcriptional Enhancers: Bridging the Genome and Phenome.Cold Spring Harb Symp Quant Biol. 2015;80:17-26. doi: 10.1101/sqb.2015.80.027219. Epub 2015 Nov 18. Cold Spring Harb Symp Quant Biol. 2015. PMID: 26582789 Review.
Cited by
-
NucHMM: a method for quantitative modeling of nucleosome organization identifying functional nucleosome states distinctly associated with splicing potentiality.Genome Biol. 2021 Aug 26;22(1):250. doi: 10.1186/s13059-021-02465-1. Genome Biol. 2021. PMID: 34446075 Free PMC article.
-
An initial genetic analysis of gemcitabine-induced high-grade neutropenia in pancreatic cancer patients in CALGB 80303 (Alliance).Pharmacogenet Genomics. 2019 Aug;29(6):123-131. doi: 10.1097/FPC.0000000000000375. Pharmacogenet Genomics. 2019. PMID: 30889042 Free PMC article. Clinical Trial.
-
A Systematic Review of Lymphangioleiomyomatosis on Diagnosis and Molecular Mechanism.Biomed Res Int. 2021 Feb 10;2021:6612776. doi: 10.1155/2021/6612776. eCollection 2021. Biomed Res Int. 2021. Retraction in: Biomed Res Int. 2023 Jul 12;2023:9823193. doi: 10.1155/2023/9823193. PMID: 33628792 Free PMC article. Retracted.
-
Progesterone Receptor Attenuates STAT1-Mediated IFN Signaling in Breast Cancer.J Immunol. 2019 May 15;202(10):3076-3086. doi: 10.4049/jimmunol.1801152. Epub 2019 Apr 1. J Immunol. 2019. PMID: 30936295 Free PMC article.
-
Hypoglycemic effect and mechanism of isoquercitrin as an inhibitor of dipeptidyl peptidase-4 in type 2 diabetic mice.RSC Adv. 2018 Apr 19;8(27):14967-14974. doi: 10.1039/c8ra00675j. eCollection 2018 Apr 18. RSC Adv. 2018. PMID: 35541310 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

