A Pathway-Centered Analysis of Pig Domestication and Breeding in Eurasia
- PMID: 28500056
- PMCID: PMC5499126
- DOI: 10.1534/g3.117.042671
A Pathway-Centered Analysis of Pig Domestication and Breeding in Eurasia
Abstract
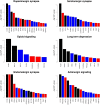
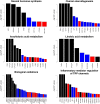
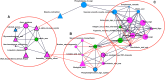
Ascertaining the molecular and physiological basis of domestication and breeding is an active area of research. Due to the current wide distribution of its wild ancestor, the wild boar, the pig (Sus scrofa) is an excellent model to study these processes, which occurred independently in East Asia and Europe ca. 9000 yr ago. Analyzing genome variability patterns in terms of metabolic pathways is attractive since it considers the impact of interrelated functions of genes, in contrast to genome-wide scans that treat genes or genome windows in isolation. To that end, we studied 40 wild boars and 123 domestic pig genomes from Asia and Europe when metabolic pathway was the unit of analysis. We computed statistical significance for differentiation (Fst) and linkage disequilibrium (nSL) statistics at the pathway level. In terms of Fst, we found 21 and 12 pathways significantly differentiated at a q-value < 0.05 in Asia and Europe, respectively; five were shared across continents. In Asia, we found six significant pathways related to behavior, which involved essential neurotransmitters like dopamine and serotonin. Several significant pathways were interrelated and shared a variable percentage of genes. There were 12 genes present in >10 significant pathways (in terms of Fst), comprising genes involved in the transduction of a large number of signals, like phospholipase PCLB1, which is expressed in the brain, or ITPR3, which has an important role in taste transduction. In terms of nSL, significant pathways were mainly related to reproductive performance (ovarian steroidogenesis), a similarly important target trait during domestication and modern animal breeding. Different levels of recombination cannot explain these results, since we found no correlation between Fst and recombination rate. However, we did find an increased ratio of deleterious mutations in domestic vs. wild populations, suggesting a relaxed functional constraint associated with the domestication and breeding processes. Purifying selection was, nevertheless, stronger in significantly differentiated pathways than in random pathways, mainly in Europe. We conclude that pathway analysis facilitates the biological interpretation of genome-wide studies. Notably, in the case of pig, behavior played an important role, among other physiological and developmental processes.
Keywords: behavior; domestication; genome sequence data; pathway analysis; pig.
Copyright © 2017 Leno-Colorado et al.
Figures
References
-
- Ai H., Fang X., Yang B., Huang Z., Chen H., et al. , 2015. Adaptation and possible ancient interspecies introgression in pigs identified by whole-genome sequencing. Nat. Genet. 47: 217–225. - PubMed
-
- Benjamini Y., Hochberg Y., 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57: 289–300.
-
- Bianco E., Soto H. W., Vargas L., Pérez-Enciso M., 2015a The chimerical genome of Isla del Coco feral pigs (Costa Rica), an isolated population since 1793 but with remarkable levels of diversity. Mol. Ecol. 24: 2364–2378. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
