The herpes viral transcription factor ICP4 forms a novel DNA recognition complex
- PMID: 28505309
- PMCID: PMC5737704
- DOI: 10.1093/nar/gkx419
The herpes viral transcription factor ICP4 forms a novel DNA recognition complex
Abstract
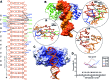
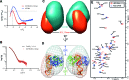
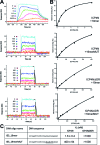
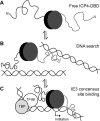
The transcription factor ICP4 from herpes simplex virus has a central role in regulating the gene expression cascade which controls viral infection. Here we present the crystal structure of the functionally essential ICP4 DNA binding domain in complex with a segment from its own promoter, revealing a novel homo-dimeric fold. We also studied the complex in solution by small angle X-Ray scattering, nuclear magnetic resonance and surface-plasmon resonance which indicated that, in addition to the globular domain, a flanking intrinsically disordered region also recognizes DNA. Together the data provides a rationale for the bi-partite nature of the ICP4 DNA recognition consensus sequence as the globular and disordered regions bind synergistically to adjacent DNA motifs. Therefore in common with its eukaryotic host, the viral transcription factor ICP4 utilizes disordered regions to enhance the affinity and tune the specificity of DNA interactions in tandem with a globular domain.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
Fold recognition insights into function of herpes ICP4 protein.Acta Biochim Pol. 2007;54(3):551-9. Epub 2007 Sep 17. Acta Biochim Pol. 2007. PMID: 17882324
-
The N terminus and C terminus of herpes simplex virus 1 ICP4 cooperate to activate viral gene expression.J Virol. 2012 Jun;86(12):6862-74. doi: 10.1128/JVI.00651-12. Epub 2012 Apr 11. J Virol. 2012. PMID: 22496239 Free PMC article.
-
Functional interaction and colocalization of the herpes simplex virus 1 major regulatory protein ICP4 with EAP, a nucleolar-ribosomal protein.Proc Natl Acad Sci U S A. 1996 May 14;93(10):4572-6. doi: 10.1073/pnas.93.10.4572. Proc Natl Acad Sci U S A. 1996. PMID: 8643445 Free PMC article.
-
A practical guide to small angle X-ray scattering (SAXS) of flexible and intrinsically disordered proteins.FEBS Lett. 2015 Sep 14;589(19 Pt A):2570-7. doi: 10.1016/j.febslet.2015.08.027. Epub 2015 Aug 29. FEBS Lett. 2015. PMID: 26320411 Review.
-
Order through disorder: The role of intrinsically disordered regions in transcription factor binding specificity.Curr Opin Struct Biol. 2021 Dec;71:110-115. doi: 10.1016/j.sbi.2021.06.011. Epub 2021 Jul 21. Curr Opin Struct Biol. 2021. PMID: 34303077 Review.
Cited by
-
Mutational pressure by host APOBEC3s more strongly affects genes expressed early in the lytic phase of herpes simplex virus-1 (HSV-1) and human polyomavirus (HPyV) infection.PLoS Pathog. 2021 Apr 30;17(4):e1009560. doi: 10.1371/journal.ppat.1009560. eCollection 2021 Apr. PLoS Pathog. 2021. PMID: 33930088 Free PMC article.
-
Small Noncoding RNA (sncRNA1) within the Latency-Associated Transcript Modulates Herpes Simplex Virus 1 Virulence and the Host Immune Response during Acute but Not Latent Infection.J Virol. 2022 Apr 13;96(7):e0005422. doi: 10.1128/jvi.00054-22. Epub 2022 Mar 7. J Virol. 2022. PMID: 35254102 Free PMC article.
-
Conserved Outer Tegument Component UL11 from Herpes Simplex Virus 1 Is an Intrinsically Disordered, RNA-Binding Protein.mBio. 2020 May 5;11(3):e00810-20. doi: 10.1128/mBio.00810-20. mBio. 2020. PMID: 32371601 Free PMC article.
-
Toward the Eradication of Herpes Simplex Virus: Vaccination and Beyond.Viruses. 2024 Sep 17;16(9):1476. doi: 10.3390/v16091476. Viruses. 2024. PMID: 39339952 Free PMC article. Review.
-
The opportunities and challenges of epigenetic approaches to manage herpes simplex infections.Expert Rev Anti Infect Ther. 2024 Dec;22(12):1123-1142. doi: 10.1080/14787210.2024.2420329. Epub 2024 Nov 6. Expert Rev Anti Infect Ther. 2024. PMID: 39466139 Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

