Prediction of NSCLC recurrence from microarray data with GEP
- PMID: 28518058
- PMCID: PMC8687152
- DOI: 10.1049/iet-syb.2016.0033
Prediction of NSCLC recurrence from microarray data with GEP
Abstract
Lung cancer is one of the deadliest diseases in the world. Non-small cell lung cancer (NSCLC) is the most common and dangerous type of lung cancer. Despite the fact that NSCLC is preventable and curable for some cases if diagnosed at early stages, the vast majority of patients are diagnosed very late. Furthermore, NSCLC usually recurs sometime after treatment. Therefore, it is of paramount importance to predict NSCLC recurrence, so that specific and suitable treatments can be sought. Nonetheless, conventional methods of predicting cancer recurrence rely solely on histopathology data and predictions are not reliable in many cases. The microarray gene expression (GE) technology provides a promising and reliable way to predict NSCLC recurrence by analysing the GE of sample cells. This study proposes a new model from GE programming to use microarray datasets for NSCLC recurrence prediction. To this end, the authors also propose a hybrid method to rank and select relevant prognostic genes that are related to NSCLC recurrence prediction. The proposed model was evaluated on real NSCLC microarray datasets and compared with other representational models. The results demonstrated the effectiveness of the proposed model.
Figures

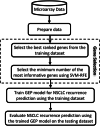





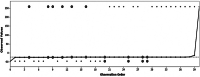
References
-
- a. c. society’ . (2015, 28/ June): Lung cancer (non‐small cell) Available at http://www.cancer.org/
-
- Lee E.‐S. Son D.‐S. Kim S.‐H. et al.: ‘Prediction of recurrence‐free survival in postoperative non‐small cell lung cancer patients by using an integrated model of clinical information and gene expression ’, Clin. Cancer Res., 2008, 14, pp. 7397–7404 - PubMed
-
- Crino L. Weder W. van Meerbeeck J. et al.: ‘Early stage and locally advanced (non‐metastatic) non‐small‐cell lung cancer: ESMO clinical practice guidelines for diagnosis, treatment and follow‐up ’, Ann. Oncol., 2010, 21, pp. v103–v115 - PubMed
-
- Hung J.‐J. Wu Y.‐C.: ‘Stage I non‐small cell lung cancer: recurrence patterns, prognostic factors and survival ’ (INTECH Open Access Publisher, 2012)
-
- Sugimura H. Nichols F.C. Yang P. et al.: ‘Survival after recurrent nonsmall‐cell lung cancer after complete pulmonary resection ’, Ann. Thorac. Surg., 2007, 83, pp. 409–418 - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

