FISH-Flow, a protocol for the concurrent detection of mRNA and protein in single cells using fluorescence in situ hybridization and flow cytometry
- PMID: 28518171
- PMCID: PMC5548662
- DOI: 10.1038/nprot.2017.039
FISH-Flow, a protocol for the concurrent detection of mRNA and protein in single cells using fluorescence in situ hybridization and flow cytometry
Abstract
We describe a flow-cytometry-based protocol for intracellular mRNA measurements in nonadherent mammalian cells using fluorescence in situ hybridization (FISH) probes. The method, which we call FISH-Flow, allows for high-throughput multiparametric measurements of gene expression, a task that was not feasible with earlier, microscopy-based approaches. The FISH-Flow protocol involves cell fixation, permeabilization and hybridization with a set of fluorescently labeled oligonucleotide probes. In this protocol, surface and intracellular protein markers can also be stained with fluorescently labeled antibodies for simultaneous protein and mRNA measurement. Moreover, a semiautomated, single-tube version of the protocol can be performed with a commercially available cell-wash device that reduces cell loss, operator time and interoperator variability. It takes ∼30 h to perform this protocol. An example of FISH-Flow measurements of cytokine mRNA induction by ex vivo stimulation of primed T cells with specific antigens is described.
Figures
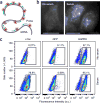




References
-
- Femino AM, Fay FS, Fogarty K, Singer RH. Visualization of single RNA transcripts in situ. Science. 1998;280:585–590. - PubMed
-
- Rahman S, Zenklusen D. Single-molecule resolution fluorescent in situ hybridization (smFISH) in the yeast S. cerevisiae. Methods Mol. Biol. 2013;1042:33–46. - PubMed
-
- Batish M, Raj A, Tyagi S. Single molecule imaging of RNA in situ. Methods Mol. Biol. 2011;714:3–13. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

