The Fragment HMGA2-sh-3p20 from HMGA2 mRNA 3'UTR Promotes the Growth of Hepatoma Cells by Upregulating HMGA2
- PMID: 28522832
- PMCID: PMC5437003
- DOI: 10.1038/s41598-017-02311-0
The Fragment HMGA2-sh-3p20 from HMGA2 mRNA 3'UTR Promotes the Growth of Hepatoma Cells by Upregulating HMGA2
Abstract
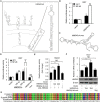
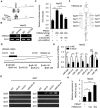
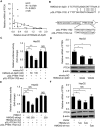
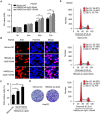
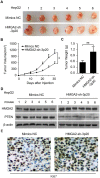
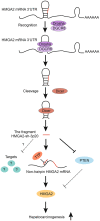
High mobility group A2 (HMGA2) plays a crucial role in the development of cancer. However, the mechanism by which HMGA2 promotes the growth of hepatocellular carcinoma (HCC) remains unclear. Here, we explore the hypothesis that HMGA2 may enhance the growth of hepatoma cells through a fragment based on the secondary structure of HMGA2 mRNA 3'-untranslated region (3'UTR). Bioinformatics analysis showed that HMGA2 mRNA displayed a hairpin structure within its 3'UTR, termed HMGA2-sh. Mechanistically, RNA immunoprecipitation assays showed that the microprocessor Drosha or DGCR8 interacted with HMGA2 mRNA in hepatoma cells. Then, Dicer contributes to the generation of the fragment HMGA2-sh-3p20 from the HMGA2-sh. HMGA2-sh-3p20 was screened by PCR analysis. Interestingly, HMGA2-sh-3p20 increased the expression of HMGA2 through antagonizing the tristetraprolin (TTP)-mediated degradation of HMGA2. HMGA2-sh-3p20 inhibited the expression of PTEN by targeting the 3'UTR of PTEN mRNA. In addition, the overexpression of PTEN could downregulate HMGA2 expression. Significantly, we documented the ability of HMGA2-sh-3p20 to promote the growth of hepatoma cells in vitro and in vivo. Thus, we conclude that the fragment HMGA2-sh-3p20 from HMGA2 mRNA 3'UTR promotes the growth of hepatoma cells by upregulating HMGA2. Our finding provides new insights into the mechanism by which HMGA2 enhances hepatocarcinogenesis.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
MiR-107 suppresses proliferation of hepatoma cells through targeting HMGA2 mRNA 3'UTR.Biochem Biophys Res Commun. 2016 Nov 18;480(3):455-460. doi: 10.1016/j.bbrc.2016.10.070. Epub 2016 Oct 20. Biochem Biophys Res Commun. 2016. PMID: 27773820
-
Post-transcriptional modulation of protein phosphatase PPP2CA and tumor suppressor PTEN by endogenous siRNA cleaved from hairpin within PTEN mRNA 3'UTR in human liver cells.Acta Pharmacol Sin. 2016 Jul;37(7):898-907. doi: 10.1038/aps.2016.18. Epub 2016 May 2. Acta Pharmacol Sin. 2016. PMID: 27133296 Free PMC article.
-
The long noncoding RNA HULC promotes liver cancer by increasing the expression of the HMGA2 oncogene via sequestration of the microRNA-186.J Biol Chem. 2017 Sep 15;292(37):15395-15407. doi: 10.1074/jbc.M117.783738. Epub 2017 Aug 1. J Biol Chem. 2017. PMID: 28765279 Free PMC article.
-
A hairpin within YAP mRNA 3'UTR functions in regulation at post-transcription level.Biochem Biophys Res Commun. 2015 Apr 3;459(2):306-312. doi: 10.1016/j.bbrc.2015.02.106. Epub 2015 Feb 26. Biochem Biophys Res Commun. 2015. PMID: 25727017
-
MiR-1297 promotes apoptosis and inhibits the proliferation and invasion of hepatocellular carcinoma cells by targeting HMGA2.Int J Mol Med. 2015 Nov;36(5):1345-52. doi: 10.3892/ijmm.2015.2341. Epub 2015 Sep 10. Int J Mol Med. 2015. PMID: 26398017
Cited by
-
Long Non-Coding RNA EGOT Promotes the Malignant Phenotypes of Hepatocellular Carcinoma Cells and Increases the Expression of HMGA2 via Down-Regulating miR-33a-5p.Onco Targets Ther. 2019 Dec 31;12:11623-11635. doi: 10.2147/OTT.S218308. eCollection 2019. Onco Targets Ther. 2019. Retraction in: Onco Targets Ther. 2022 Oct 05;15:1121-1122. doi: 10.2147/OTT.S392090. PMID: 32021242 Free PMC article. Retracted.
-
Roles of Tristetraprolin in Tumorigenesis.Int J Mol Sci. 2018 Oct 29;19(11):3384. doi: 10.3390/ijms19113384. Int J Mol Sci. 2018. PMID: 30380668 Free PMC article. Review.
-
Tumor-Linked Macrophages Promote HCC Development by Mediating the CCAT1/Let-7b/HMGA2 Signaling Pathway.Onco Targets Ther. 2020 Dec 14;13:12829-12843. doi: 10.2147/OTT.S283786. eCollection 2020. Onco Targets Ther. 2020. PMID: 33363387 Free PMC article.
-
ZFP36 Inhibits Tumor Progression of Human Prostate Cancer by Targeting CDK6 and Oxidative Stress.Oxid Med Cell Longev. 2022 Sep 6;2022:3611540. doi: 10.1155/2022/3611540. eCollection 2022. Oxid Med Cell Longev. 2022. Retraction in: Oxid Med Cell Longev. 2023 Jun 21;2023:9865647. doi: 10.1155/2023/9865647. PMID: 36111167 Free PMC article. Retracted.
-
The role of RNA-binding protein tristetraprolin in cancer and immunity.Med Oncol. 2017 Nov 9;34(12):196. doi: 10.1007/s12032-017-1055-6. Med Oncol. 2017. PMID: 29124478 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

