Musashi-1 promotes a cancer stem cell lineage and chemoresistance in colorectal cancer cells
- PMID: 28526879
- PMCID: PMC5438397
- DOI: 10.1038/s41598-017-02057-9
Musashi-1 promotes a cancer stem cell lineage and chemoresistance in colorectal cancer cells
Abstract
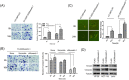
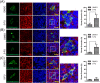
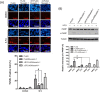
Colorectal cancers (CRCs) are a critical health issue worldwide. Cancer stem cell (CSC) lineages are associated with tumour transformation, progression, and malignant transformation. However, how lineages are transformed and how chemoresistance is acquired by CRCs remain largely unknown. In this report, we demonstrated that the RNA-binding protein Musashi-1 enhanced the development of CD44+ colorectal CSCs and triggered the formation of anti-apoptotic stress granules (SGs). Our results indicated that CD44+ CSC lineage-specific induction of tumour malignancies was controlled by Musashi-1. In addition, Musashi-1 formed SGs when CRC cell lines were treated with 5-fluorouracil. The C-terminal domain of Musashi-1 was critical for recruitment of Musashi-1 into SGs. Intracellular Musashi-1 SGs enhanced the chemoresistance of CRCs. Analysis of clinical CRC samples indicated that Musashi-1 expression was prominent in CRC stage IIA and IIB. In summary, we demonstrated that Musashi-1, a stemness gene, is a critical modulator that promotes the development of CD44+ colorectal CSCs and also enhances CRC chemoresistance via formation of SGs. Our findings elucidated a novel mechanism of CRC chemoresistance through increased anti-apoptotic effects via Musashi-1-associated SGs.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





Similar articles
-
TFAP2C promotes stemness and chemotherapeutic resistance in colorectal cancer via inactivating hippo signaling pathway.J Exp Clin Cancer Res. 2018 Feb 13;37(1):27. doi: 10.1186/s13046-018-0683-9. J Exp Clin Cancer Res. 2018. PMID: 29439714 Free PMC article.
-
PRDM14 promotes malignant phenotype and correlates with poor prognosis in colorectal cancer.Clin Transl Oncol. 2020 Jul;22(7):1126-1137. doi: 10.1007/s12094-019-02239-z. Epub 2019 Nov 18. Clin Transl Oncol. 2020. PMID: 31741141
-
Musashi-1 promotes chemoresistant granule formation by PKR/eIF2α signalling cascade in refractory glioblastoma.Biochim Biophys Acta Mol Basis Dis. 2018 May;1864(5 Pt A):1850-1861. doi: 10.1016/j.bbadis.2018.02.017. Epub 2018 Feb 24. Biochim Biophys Acta Mol Basis Dis. 2018. PMID: 29486283
-
MicroRNAs and colorectal cancer chemoresistance: New solution for old problem.Life Sci. 2020 Oct 15;259:118255. doi: 10.1016/j.lfs.2020.118255. Epub 2020 Aug 17. Life Sci. 2020. PMID: 32818543 Review.
-
Musashi RNA-Binding Proteins as Cancer Drivers and Novel Therapeutic Targets.Clin Cancer Res. 2017 May 1;23(9):2143-2153. doi: 10.1158/1078-0432.CCR-16-2728. Epub 2017 Jan 31. Clin Cancer Res. 2017. PMID: 28143872 Free PMC article. Review.
Cited by
-
Musashi-1 Is a Novel Immunohistochemical Marker of Neuroendocrine Carcinoma of the Lung.Cancers (Basel). 2023 Nov 29;15(23):5631. doi: 10.3390/cancers15235631. Cancers (Basel). 2023. PMID: 38067335 Free PMC article.
-
Musashi-1: An Example of How Polyalanine Tracts Contribute to Self-Association in the Intrinsically Disordered Regions of RNA-Binding Proteins.Int J Mol Sci. 2020 Mar 26;21(7):2289. doi: 10.3390/ijms21072289. Int J Mol Sci. 2020. PMID: 32225071 Free PMC article.
-
Stress granules and hormetic adaptation of cancer.Trends Cancer. 2023 Dec;9(12):995-1005. doi: 10.1016/j.trecan.2023.08.005. Epub 2023 Sep 11. Trends Cancer. 2023. PMID: 37704502 Free PMC article. Review.
-
RNA Binding Proteins as Potential Therapeutic Targets in Colorectal Cancer.Cancers (Basel). 2024 Oct 16;16(20):3502. doi: 10.3390/cancers16203502. Cancers (Basel). 2024. PMID: 39456596 Free PMC article. Review.
-
Upregulation of musashi1 increases malignancy of hepatocellular carcinoma via the Wnt/β-catenin signaling pathway and predicts a poor prognosis.BMC Gastroenterol. 2019 Dec 30;19(1):230. doi: 10.1186/s12876-019-1150-6. BMC Gastroenterol. 2019. PMID: 31888604 Free PMC article.
References
-
- Heinen CD, Shivapurkar N, Tang Z, Groden J, Alabaster O. Microsatellite instability in aberrant crypt foci from human colons. Cancer research. 1996;56:5339–5341. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

