Applications of NMR to membrane proteins
- PMID: 28529197
- PMCID: PMC5657258
- DOI: 10.1016/j.abb.2017.05.011
Applications of NMR to membrane proteins
Abstract
Membrane proteins present a challenge for structural biology. In this article, we review some of the recent developments that advance the application of NMR to membrane proteins, with emphasis on structural studies in detergent-free, lipid bilayer samples that resemble the native environment. NMR spectroscopy is not only ideally suited for structure determination of membrane proteins in hydrated lipid bilayer membranes, but also highly complementary to the other principal techniques based on X-ray and electron diffraction. Recent advances in NMR instrumentation, spectroscopic methods, computational methods, and sample preparations are driving exciting new efforts in membrane protein structural biology.
Keywords: Bilayer; Lipid; Membrane protein; NMR; Solid-state NMR.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures
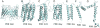
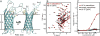
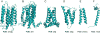
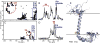
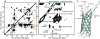

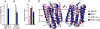
Similar articles
-
Applications of solid-state NMR to membrane proteins.Biochim Biophys Acta Proteins Proteom. 2017 Nov;1865(11 Pt B):1577-1586. doi: 10.1016/j.bbapap.2017.07.004. Epub 2017 Jul 12. Biochim Biophys Acta Proteins Proteom. 2017. PMID: 28709996 Review.
-
Recent advances in magic angle spinning solid state NMR of membrane proteins.Prog Nucl Magn Reson Spectrosc. 2014 Oct;82:1-26. doi: 10.1016/j.pnmrs.2014.07.001. Epub 2014 Jul 26. Prog Nucl Magn Reson Spectrosc. 2014. PMID: 25444696 Review.
-
Solid-state NMR structures of integral membrane proteins.Mol Membr Biol. 2015 Aug-Dec;32(5-8):156-78. doi: 10.3109/09687688.2016.1139754. Epub 2016 Feb 8. Mol Membr Biol. 2015. PMID: 26857803 Review.
-
Detergent optimized membrane protein reconstitution in liposomes for solid state NMR.Biochemistry. 2014 Apr 22;53(15):2454-63. doi: 10.1021/bi500144h. Epub 2014 Apr 9. Biochemistry. 2014. PMID: 24665863 Free PMC article.
-
Synthetic Biology-Based Solution NMR Studies on Membrane Proteins in Lipid Environments.Methods Enzymol. 2019;614:143-185. doi: 10.1016/bs.mie.2018.08.019. Epub 2018 Dec 21. Methods Enzymol. 2019. PMID: 30611423
Cited by
-
Improved Protocol for the Production of the Low-Expression Eukaryotic Membrane Protein Human Aquaporin 2 in Pichia pastoris for Solid-State NMR.Biomolecules. 2020 Mar 11;10(3):434. doi: 10.3390/biom10030434. Biomolecules. 2020. PMID: 32168846 Free PMC article.
-
Dimer Organization of Membrane-Associated NS5A of Hepatitis C Virus as Determined by Highly Sensitive 1 H-Detected Solid-State NMR.Angew Chem Int Ed Engl. 2021 Mar 1;60(10):5339-5347. doi: 10.1002/anie.202013296. Epub 2021 Jan 18. Angew Chem Int Ed Engl. 2021. PMID: 33205864 Free PMC article.
-
The Lassa Virus Stable Signal Peptide Undergoes a Conformational Change to Aid Viral Fusion.Chemistry. 2025 Mar 25;31(18):e202403608. doi: 10.1002/chem.202403608. Epub 2025 Mar 7. Chemistry. 2025. PMID: 39946735 Free PMC article.
-
Dipolar Recoupling in Rotating Solids.Chem Rev. 2024 Nov 27;124(22):12844-12917. doi: 10.1021/acs.chemrev.4c00373. Epub 2024 Nov 6. Chem Rev. 2024. PMID: 39504237 Free PMC article. Review.
-
Formation of the β-barrel assembly machinery complex in lipid bilayers as seen by solid-state NMR.Nat Commun. 2018 Oct 8;9(1):4135. doi: 10.1038/s41467-018-06466-w. Nat Commun. 2018. PMID: 30297837 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

