Enhancing poly-γ-glutamic acid production in Bacillus amyloliquefaciens by introducing the glutamate synthesis features from Corynebacterium glutamicum
- PMID: 28532451
- PMCID: PMC5440981
- DOI: 10.1186/s12934-017-0704-y
Enhancing poly-γ-glutamic acid production in Bacillus amyloliquefaciens by introducing the glutamate synthesis features from Corynebacterium glutamicum
Abstract
Background: Poly-γ-glutamic acid (γ-PGA) is a valuable polymer with glutamate as its sole precursor. Enhancement of the intracellular glutamate synthesis is a very important strategy for the improvement of γ-PGA production, especially for those glutamate-independent γ-PGA producing strains. Corynebacterium glutamicum has long been used for industrial glutamate production and it exhibits some unique features for glutamate synthesis; therefore introduction of these metabolic characters into the γ-PGA producing strain might lead to increased intracellular glutamate availability, and thus ultimate γ-PGA production.
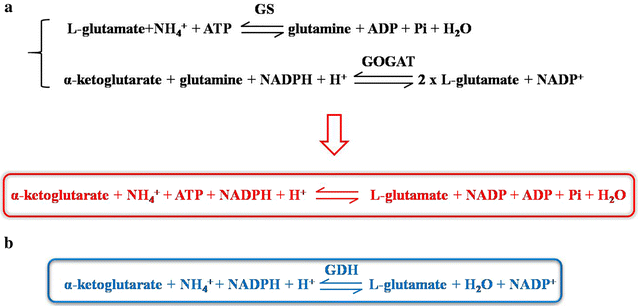
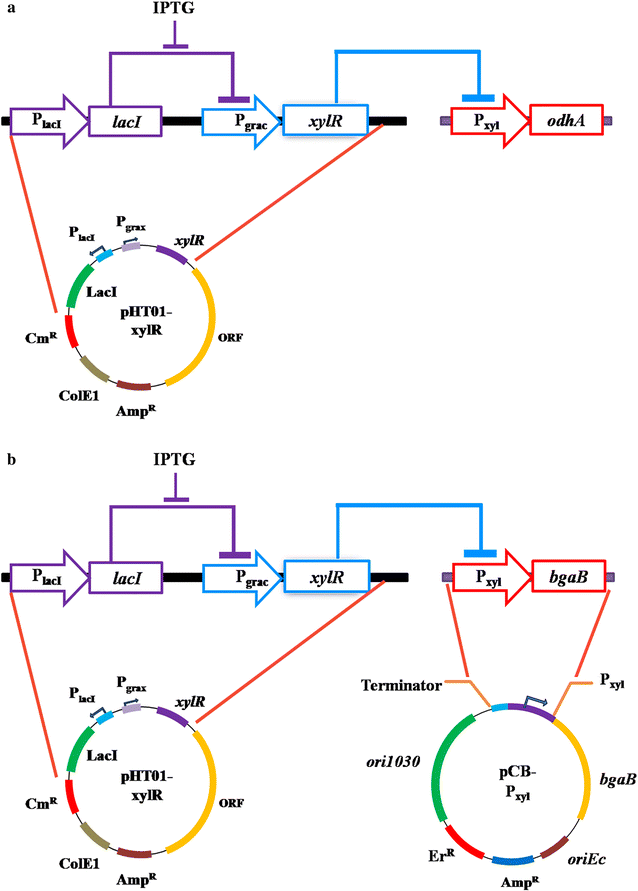
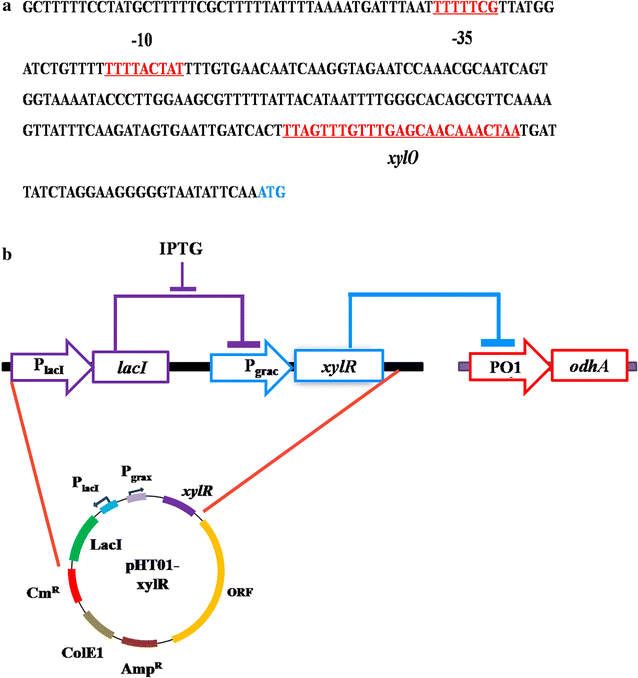
Results: In this study, the unique glutamate synthesis features from C. glutamicum was introduced into the glutamate-independent γ-PGA producing Bacillus amyloliquefaciens NK-1 strain. After introducing the energy-saving NADPH-dependent glutamate dehydrogenase (NADPH-GDH) pathway, the NK-1 (pHT315-gdh) strain showed slightly increase (by 9.1%) in γ-PGA production. Moreover, an optimized metabolic toggle switch for controlling the expression of ɑ-oxoglutarate dehydrogenase complex (ODHC) was introduced into the NK-1 strain, because it was previously shown that the ODHC in C. glutamicum was completely inhibited when glutamate was actively produced. The obtained NK-PO1 (pHT01-xylR) strain showed 66.2% higher γ-PGA production than the NK-1 strain. However, the further combination of these two strategies (introducing both NADPH-GDH pathway and the metabolic toggle switch) did not lead to further increase of γ-PGA production but rather the resultant γ-PGA production was even lower than that in the NK-1 strain.
Conclusions: We proposed new metabolic engineering strategies to improve the γ-PGA production in B. amyloliquefaciens. The NK-1 (pHT315-gdh) strain with the introduction of NADPH-GDH pathway showed 9.1% improvement in γ-PGA production. The NK-PO1 (pHT01-xylR) strain with the introduction of a metabolic toggle switch for controlling the expression of ODHC showed 66.2% higher γ-PGA production than the NK-1 strain. This work proposed a new strategy for improving the target product in microbial cell factories.
Keywords: Metabolic toggle switch; NADPH-dependent glutamate dehydrogenase; Poly-γ-glutamic acid.
Figures



Similar articles
-
Genetic and metabolic engineering for poly-γ-glutamic acid production: current progress, challenges, and prospects.World J Microbiol Biotechnol. 2022 Aug 28;38(11):208. doi: 10.1007/s11274-022-03390-6. World J Microbiol Biotechnol. 2022. PMID: 36030456 Review.
-
Metabolic engineering of Bacillus amyloliquefaciens LL3 for enhanced poly-γ-glutamic acid synthesis.Microb Biotechnol. 2019 Sep;12(5):932-945. doi: 10.1111/1751-7915.13446. Epub 2019 Jun 20. Microb Biotechnol. 2019. PMID: 31219230 Free PMC article.
-
Improved poly-γ-glutamic acid production in Bacillus amyloliquefaciens by modular pathway engineering.Metab Eng. 2015 Nov;32:106-115. doi: 10.1016/j.ymben.2015.09.011. Epub 2015 Sep 26. Metab Eng. 2015. PMID: 26410449
-
Enhancing Poly-γ-glutamic Acid Production in Bacillus tequilensis BL01 through a Multienzyme Assembly Strategy and Expression Features of Glutamate Synthesis from Corynebacterium glutamicum.J Agric Food Chem. 2024 Apr 17;72(15):8674-8683. doi: 10.1021/acs.jafc.4c00685. Epub 2024 Apr 3. J Agric Food Chem. 2024. PMID: 38569079
-
The Actinobacterium Corynebacterium glutamicum, an Industrial Workhorse.J Microbiol Biotechnol. 2016 May 28;26(5):807-22. doi: 10.4014/jmb.1601.01053. J Microbiol Biotechnol. 2016. PMID: 26838341 Review.
Cited by
-
Genetic and metabolic engineering for poly-γ-glutamic acid production: current progress, challenges, and prospects.World J Microbiol Biotechnol. 2022 Aug 28;38(11):208. doi: 10.1007/s11274-022-03390-6. World J Microbiol Biotechnol. 2022. PMID: 36030456 Review.
-
Metabolic engineering of Bacillus amyloliquefaciens LL3 for enhanced poly-γ-glutamic acid synthesis.Microb Biotechnol. 2019 Sep;12(5):932-945. doi: 10.1111/1751-7915.13446. Epub 2019 Jun 20. Microb Biotechnol. 2019. PMID: 31219230 Free PMC article.
-
Enhanced production of poly-γ-glutamic acid via optimizing the expression cassette of Vitreoscilla hemoglobin in Bacillus licheniformis.Synth Syst Biotechnol. 2022 Jan 27;7(1):567-573. doi: 10.1016/j.synbio.2022.01.006. eCollection 2022 Mar. Synth Syst Biotechnol. 2022. PMID: 35155838 Free PMC article.
-
Metabolic Engineering of Central Carbon Metabolism of Bacillus licheniformis for Enhanced Production of Poly-γ-glutamic Acid.Appl Biochem Biotechnol. 2021 Nov;193(11):3540-3552. doi: 10.1007/s12010-021-03619-4. Epub 2021 Jul 26. Appl Biochem Biotechnol. 2021. PMID: 34312784
-
Modular Engineering to Enhance Keratinase Production for Biotransformation of Discarded Feathers.Appl Biochem Biotechnol. 2023 Mar;195(3):1752-1769. doi: 10.1007/s12010-022-04206-x. Epub 2022 Nov 17. Appl Biochem Biotechnol. 2023. PMID: 36394712
References
-
- Ashiuchi M, Misono H. Biochemistry and molecular genetics of poly-γ-glutamate synthesis. Appl Biochem Biotechnol. 2002;59:9–14. - PubMed
-
- Kunioka M. Biosynthesis and chemical reactions of poly(amino acid)s from microorganisms. Appl Microbiol Biotechnol. 1997;47:469–475. doi: 10.1007/s002530050958. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

