Essential role for histone deacetylase 11 (HDAC11) in neutrophil biology
- PMID: 28550123
- PMCID: PMC6608079
- DOI: 10.1189/jlb.1A0415-176RRR
Essential role for histone deacetylase 11 (HDAC11) in neutrophil biology
Abstract
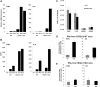
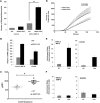
Epigenetic changes in chromatin structure have been recently associated with the deregulated expression of critical genes in normal and malignant processes. HDAC11, the newest member of the HDAC family of enzymes, functions as a negative regulator of IL-10 expression in APCs, as previously described by our lab. However, at the present time, its role in other hematopoietic cells, specifically in neutrophils, has not been fully explored. In this report, for the first time, we present a novel physiologic role for HDAC11 as a multifaceted regulator of neutrophils. Thus far, we have been able to demonstrate a lineage-restricted overexpression of HDAC11 in neutrophils and committed neutrophil precursors (promyelocytes). Additionally, we show that HDAC11 appears to associate with the transcription machinery, possibly regulating the expression of inflammatory and migratory genes in neutrophils. Given the prevalence of neutrophils in the peripheral circulation and their central role in the first line of defense, our results highlight a unique and novel role for HDAC11. With the consideration of the emergence of new, selective HDAC11 inhibitors, we believe that our findings will have significant implications in a wide range of diseases spanning malignancies, autoimmunity, and inflammation.
Keywords: epigenetics; gene regulation; innate immunity.
© Society for Leukocyte Biology.
Figures




References
-
- Nicolás‐Ávila, J. A. , Adrover, J. M. , Hidalgo, A. (2017) Neutrophils in homeostasis, immunity, and cancer. Immunity 46, 15–28. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

