Inflammation-associated DNA methylation patterns in epithelium of ulcerative colitis
- PMID: 28557546
- PMCID: PMC5687324
- DOI: 10.1080/15592294.2017.1334023
Inflammation-associated DNA methylation patterns in epithelium of ulcerative colitis
Abstract
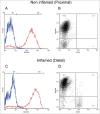
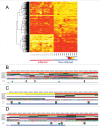
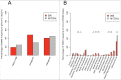
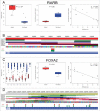
Aberrant DNA methylation patterns have been reported in inflamed tissues and may play a role in disease. We studied DNA methylation and gene expression profiles of purified intestinal epithelial cells from ulcerative colitis patients, comparing inflamed and non-inflamed areas of the colon. We identified 577 differentially methylated sites (false discovery rate <0.2) mapping to 210 genes. From gene expression data from the same epithelial cells, we identified 62 differentially expressed genes with increased expression in the presence of inflammation at prostate cancer susceptibility genes PRAC1 and PRAC2. Four genes showed inverse correlation between methylation and gene expression; ROR1, GXYLT2, FOXA2, and, notably, RARB, a gene previously identified as a tumor suppressor in colorectal adenocarcinoma as well as breast, lung and prostate cancer. We highlight targeted and specific patterns of DNA methylation and gene expression in epithelial cells from inflamed colon, while challenging the importance of epithelial cells in the pathogenesis of chronic inflammation.
Keywords: DNA methylation; inflammatory bowel disease; intestinal epithelial cell; transcriptome; ulcerative colitis.
Figures

References
-
- Conrad K, Roggenbuck D, Laass MW. Diagnosis and classification of ulcerative colitis. Autoimmun Rev 2014; 13:463–6; PMID:24424198; https://doi.org/10.1016/j.autrev.2014.01.028 - DOI - PubMed
-
- Laass MW, Roggenbuck D, Conrad K. Diagnosis and classification of Crohn's disease. Autoimmun Rev 13:467–71; PMID:24424189; https://doi.org/10.1016/j.autrev.2014.01.029</bib> - DOI - PubMed
-
- Li Y, Kundu P, Seow SW, de Matos CT, Aronsson L, Chin KC, Kärre K, Pettersson S, Greicius G. Gut microbiota accelerate tumor growth via c-jun and STAT3 phosphorylation in APCMin/+ mice. Carcinogenesis 2012; 33:1231–8; PMID:22461519; https://doi.org/10.1093/carcin/bgs137 - DOI - PubMed
-
- Jostins L, Ripke S, Weersma RK, Duerr RH, McGovern DP, Hui KY, Lee JC, Schumm LP, Sharma Y, Anderson CA, et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 2012; 491:119–24; PMID:23128233; https://doi.org/10.1038/nature11582 - DOI - PMC - PubMed
-
- Araki A, Kanai T, Ishikura T, Makita S, Uraushihara K, Iiyama R, Totsuka T, Takeda K, Akira S, Watanabe M. MyD88-deficient mice develop severe intestinal inflammation in dextran sodium sulfate colitis. J Gastroenterol 2005; 40:16–23; PMID:15692785; https://doi.org/10.1007/s00535-004-1492-9 - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
