Buffers Strongly Modulate Fibrin Self-Assembly into Fibrous Networks
- PMID: 28558246
- PMCID: PMC5489959
- DOI: 10.1021/acs.langmuir.7b00527
Buffers Strongly Modulate Fibrin Self-Assembly into Fibrous Networks
Abstract
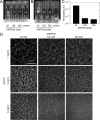
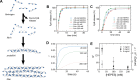
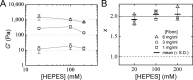
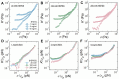
Fibrin is a plasma protein with a central role in blood clotting and wound repair. Upon vascular injury, fibrin forms resilient fibrillar networks (clots) via a multistep self-assembly process, from monomers, to double-stranded protofibrils, to a branched network of thick fibers. In vitro, fibrin self-assembly is sensitive to physicochemical conditions like the solution pH and ionic strength, which tune the strength of the noncovalent driving forces. Here we report a surprising finding that the buffer-which is necessary to control the pH and is typically considered to be inert-also significantly influences fibrin self-assembly. We show by confocal microscopy and quantitative light scattering that various common buffering agents have no effect on the initial assembly of fibrin monomers into protofibrils but strongly hamper the subsequent lateral association of protofibrils into thicker fibers. We further find that the structural changes are independent of the molecular structure of the buffering agents as well as of the activation mechanism and even occur in fibrin networks formed from platelet-poor plasma. This buffer-mediated decrease in protofibril bundling results in a marked reduction in the permeability of fibrin networks but only weakly influences the elastic modulus of fibrin networks, providing a useful tuning parameter to independently control the elastic properties and the permeability of fibrin networks. Our work raises the possibility that fibrin assembly in vivo may be regulated by variations in the acute-phase levels of bicarbonate and phosphate, which act as physiological buffering agents of blood pH. Moreover, our findings add a new example of buffer-induced effects on biomolecular self-assembly to recent findings for a range of proteins and lipids.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


Similar articles
-
Fibrin self-assembly is adapted to oxidation.Free Radic Biol Med. 2016 Jun;95:55-64. doi: 10.1016/j.freeradbiomed.2016.03.005. Epub 2016 Mar 9. Free Radic Biol Med. 2016. PMID: 26969792
-
Assembly of fibrin. A light scattering study.J Biol Chem. 1979 Nov 25;254(22):11272-81. J Biol Chem. 1979. PMID: 500644
-
Does topology drive fiber polymerization?Biochemistry. 2014 Dec 16;53(49):7824-34. doi: 10.1021/bi500986z. Epub 2014 Dec 5. Biochemistry. 2014. PMID: 25419972 Free PMC article.
-
Fibrinogen and fibrin.Adv Protein Chem. 2005;70:247-99. doi: 10.1016/S0065-3233(05)70008-5. Adv Protein Chem. 2005. PMID: 15837518 Review.
-
Thrombin generation and fibrin clot structure.Blood Rev. 2007 May;21(3):131-42. doi: 10.1016/j.blre.2006.11.001. Epub 2007 Jan 8. Blood Rev. 2007. PMID: 17208341 Review.
Cited by
-
Increasing salinity of fibrinogen solvent generates stable fibrin hydrogels for cell delivery or tissue engineering.PLoS One. 2021 May 19;16(5):e0239242. doi: 10.1371/journal.pone.0239242. eCollection 2021. PLoS One. 2021. PMID: 34010323 Free PMC article.
-
Proteomic Evidence for Amyloidogenic Cross-Seeding in Fibrinaloid Microclots.Int J Mol Sci. 2024 Oct 8;25(19):10809. doi: 10.3390/ijms251910809. Int J Mol Sci. 2024. PMID: 39409138 Free PMC article.
-
Snake venom-defined fibrin architecture dictates fibroblast survival and differentiation.Nat Commun. 2023 Feb 23;14(1):1029. doi: 10.1038/s41467-023-36437-9. Nat Commun. 2023. PMID: 36823141 Free PMC article.
-
Microrheology and structural quantification of hypercoagulable clots.Biomed Opt Express. 2023 Jul 19;14(8):4179-4189. doi: 10.1364/BOE.492669. eCollection 2023 Aug 1. Biomed Opt Express. 2023. PMID: 37799698 Free PMC article.
-
Laser speckle rheological microscopy reveals wideband viscoelastic spectra of biological tissues.Sci Adv. 2024 May 10;10(19):eadl1586. doi: 10.1126/sciadv.adl1586. Epub 2024 May 8. Sci Adv. 2024. PMID: 38718128 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

