Spatial regulation of coalesced protein assemblies: Lessons from yeast to diseases
- PMID: 28574744
- PMCID: PMC5480387
- DOI: 10.1080/19336896.2017.1322239
Spatial regulation of coalesced protein assemblies: Lessons from yeast to diseases
Abstract
Organisms rely on correctly folded proteins to carry out essential functions. Protein quality control factors guard proteostasis and prevent protein misfolding. When quality control fails and in response to diverse stresses, many proteins start to accumulate at specific deposit sites that maintain cellular organization and protect the functionality of coalescing proteins. These transitions involve dedicated proteins that promote coalescence and are facilitated by endo-membranes and cytoskeletal platforms. Moreover, several proteins make use of weak multivalent interactions or conformational templating to drive the formation of large-scale assemblies. Formation of such assemblies is often associated with a change in biochemical activity that can be used by cells to execute biochemical decisions in a localized manner during development and adaption. Since all assembly types impact cell physiology, their localization and dynamics need to be tightly regulated. Interestingly, at least some of the regulatory mechanisms are shared by functional membrane-less organelles and assemblies of terminally aggregated proteins. Furthermore, constituents of functional assemblies can aggregate and become non-functional during aging. Here we present the current knowledge as to how coalescing protein assemblies are spatially organized in cells and we postulate that failures in their spatial confinement might underscore certain aspects of aging and neurodegenerative diseases.
Figures
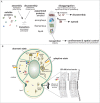
Comment on
- Extra View to: Saarikangas J, Caudron F, Prasad R, Moreno DF, Bolognesi A, Aldea M, Barral Y. Compartmentalization of ER-bound chaperone confines protein deposit formation to the aging yeast cell. Curr Biol 2017; 27:773–83; https://doi.org/10.1016/j.cub.2017.01.069
Similar articles
-
Macromolecular complexes as depots for releasable regulatory proteins.Trends Biochem Sci. 2007 Apr;32(4):158-64. doi: 10.1016/j.tibs.2007.02.003. Epub 2007 Feb 23. Trends Biochem Sci. 2007. PMID: 17321138 Review.
-
Mechanisms Underlying the Essential Role of Mitochondrial Membrane Lipids in Yeast Chronological Aging.Oxid Med Cell Longev. 2017;2017:2916985. doi: 10.1155/2017/2916985. Epub 2017 May 16. Oxid Med Cell Longev. 2017. PMID: 28593023 Free PMC article. Review.
-
Absence of Non-histone Protein Complexes at Natural Chromosomal Pause Sites Results in Reduced Replication Pausing in Aging Yeast Cells.Cell Rep. 2016 Nov 8;17(7):1747-1754. doi: 10.1016/j.celrep.2016.10.050. Cell Rep. 2016. PMID: 27829146 Free PMC article.
-
The Intrinsically Disordered Protein Atg13 Mediates Supramolecular Assembly of Autophagy Initiation Complexes.Dev Cell. 2016 Jul 11;38(1):86-99. doi: 10.1016/j.devcel.2016.06.015. Dev Cell. 2016. PMID: 27404361
-
The multifaceted roles of intrinsic disorder in protein complexes.FEBS Lett. 2015 Sep 14;589(19 Pt A):2498-506. doi: 10.1016/j.febslet.2015.06.004. Epub 2015 Jun 11. FEBS Lett. 2015. PMID: 26073257 Review.
References
-
- Saarikangas J, Barral Y. Protein aggregation as a mechanism of adaptive cellular responses. Curr Genet 2016; 62:711-24; PMID:27032776; https//doi.org/10.1007/s00294-016-0596-0 - DOI - PubMed
-
- Aguzzi A, Lakkaraju AKK. Cell biology of Prions and Prionoids: A status report. Trends Cell Biol 2016; 26:40-51; PMID:26455408; https//doi.org/10.1016/j.tcb.2015.08.007 - DOI - PubMed
-
- Wallace EWJ, Kear-Scott JL, Pilipenko EV, Schwartz MH, Laskowski PR, Rojek AE, Katanski CD, Riback JA, Dion MF, Franks AM, et al.. Reversible, specific, active aggregates of endogenous proteins assemble upon heat stress. Cell 2015; 162:1286-98; PMID:26359986; https//doi.org/10.1016/j.cell.2015.08.041 - DOI - PMC - PubMed
-
- Caudron F, Barral Y. A super-assembly of Whi3 encodes memory of deceptive encounters by single cells during yeast courtship. Cell 2013; 155:1244-57; PMID:24315096; https//doi.org/10.1016/j.cell.2013.10.046 - DOI - PubMed
-
- Li L, Sanchez CP, Slaughter BD, Zhao Y, Khan MR, Unruh JR, Rubinstein B, Si K. A putative biochemical engram of long-term memory. Curr Biol 2016; 26(23):3143-56; PMID:27818176; https//doi.org/10.1016/j.cub.2016.09.054 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
