Tension sensors reveal how the kinetochore shares its load
- PMID: 28582586
- PMCID: PMC5543933
- DOI: 10.1002/bies.201600216
Tension sensors reveal how the kinetochore shares its load
Abstract
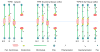
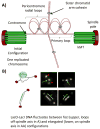
At metaphase in mitotic cells, pulling forces at the kinetochore-microtubule interface create tension by stretching the centromeric chromatin between oppositely oriented sister kinetochores. This tension is important for stabilizing the end-on kinetochore microtubule attachment required for proper bi-orientation of sister chromosomes as well as for satisfaction of the Spindle Assembly Checkpoint and entry into anaphase. How force is coupled by proteins to kinetochore microtubules and resisted by centromere stretch is becoming better understood as many of the proteins involved have been identified. Recent application of genetically encoded fluorescent tension sensors within the mechanical linkage between the centromere and kinetochore microtubules are beginning to reveal - from live cell assays - protein specific contributions that are functionally important.
Keywords: centromeric chromatin; chromosome segregation; kinetochore; microtubules; mitosis; tension sensor.
© 2017 WILEY Periodicals, Inc.
Figures



References
-
- Hori T, Amano M, Suzuki A, Backer CB, et al. CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell. 2008;135:1039–52. - PubMed
-
- Pekgoz Altunkaya G, Malvezzi F, Demianova Z, Zimniak T, et al. CCAN assembly configures composite binding interfaces to promote cross-linking of Ndc80 complexes at the kinetochore. Curr Biol. 2016;26:2370–8. - PubMed
-
- Przewloka MR, Venkei Z, Bolanos-Garcia VM, Debski J, et al. CENP-C is a structural platform for kinetochore assembly. Curr Biol. 2011;21:399–405. - PubMed
-
- Schleiffer A, Maier M, Litos G, Lampert F, et al. CENP-T proteins are conserved centromere receptors of the Ndc80 complex. Nat Cell Biol. 2012;14:604–13. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

