MYC Deregulation in Primary Human Cancers
- PMID: 28587062
- PMCID: PMC5485515
- DOI: 10.3390/genes8060151
MYC Deregulation in Primary Human Cancers
Abstract
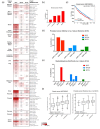
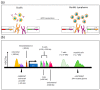
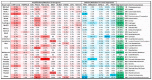
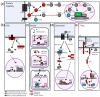
MYC regulates a complex biological program by transcriptionally activating and repressing its numerous target genes. As such, MYC is a master regulator of many processes, including cell cycle entry, ribosome biogenesis, and metabolism. In cancer, the activity of the MYC transcriptional network is frequently deregulated, contributing to the initiation and maintenance of disease. Deregulation often leads to constitutive overexpression of MYC, which can be achieved through gross genetic abnormalities, including copy number alterations, chromosomal translocations, increased enhancer activity, or through aberrant signal transduction leading to increased MYC transcription or increased MYC mRNA and protein stability. Herein, we summarize the frequency and modes of MYC deregulation and describe both well-established and more recent findings in a variety of cancer types. Notably, these studies have highlighted that with an increased appreciation for the basic mechanisms deregulating MYC in cancer, new therapeutic vulnerabilities can be discovered and potentially exploited for the inhibition of this potent oncogene in cancer.
Keywords: MYC; cancer; cell signaling; deregulation; enhancers; gene amplification; translocation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Mao D.Y., Watson J.D., Yan P.S., Barsyte-Lovejoy D., Khosravi F., Wong W.W., Farnham P.J., Huang T.H., Penn L.Z. Analysis of MYC bound loci identified by CpG island arrays shows that MAX is essential for MYC-dependent repression. Curr. Biol. 2003;13:882–886. doi: 10.1016/S0960-9822(03)00297-5. - DOI - PubMed
-
- Mateyak M.K., Obaya A.J., Adachi S., Sedivy J.M. Phenotypes of c-MYC-deficient rat fibroblasts isolated by targeted homologous recombination. Cell Growth Differ. 1997;8:1039–1048. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

