Two Glycerol-3-Phosphate Dehydrogenases from Chlamydomonas Have Distinct Roles in Lipid Metabolism
- PMID: 28588114
- PMCID: PMC5543956
- DOI: 10.1104/pp.17.00491
Two Glycerol-3-Phosphate Dehydrogenases from Chlamydomonas Have Distinct Roles in Lipid Metabolism
Abstract
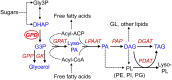
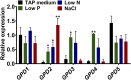
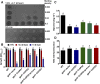
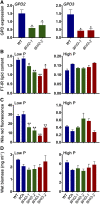
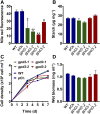
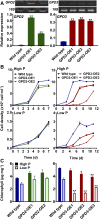
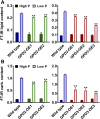
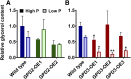
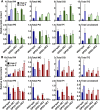
The metabolism of glycerol-3-phosphate (G3P) is important for environmental stress responses by eukaryotic microalgae. G3P is an essential precursor for glycerolipid synthesis and the accumulation of triacylglycerol (TAG) in response to nutrient starvation. G3P dehydrogenase (GPDH) mediates G3P synthesis, but the roles of specific GPDH isoforms are currently poorly understood. Of the five GPDH enzymes in the model alga Chlamydomonas reinhardtii, GPD2 and GPD3 were shown to be induced by nutrient starvation and/or salt stress. Heterologous expression of GPD2, a putative chloroplastic GPDH, and GPD3, a putative cytosolic GPDH, in a yeast gpd1Δ mutant demonstrated the functionality of both enzymes. C. reinhardtii knockdown mutants for GPD2 and GPD3 showed no difference in growth but displayed significant reduction in TAG concentration compared with the wild type in response to phosphorus or nitrogen starvation. Overexpression of GPD2 and GPD3 in C. reinhardtii gave distinct phenotypes. GPD2 overexpression lines showed only subtle metabolic phenotypes and no significant alteration in growth. In contrast, GPD3 overexpression lines displayed significantly inhibited growth and chlorophyll concentration, reduced glycerol concentration, and changes to lipid composition compared with the wild type, including increased abundance of phosphatidic acids but reduced abundance of diglycerides, triglycerides, and phosphatidylglycerol lipids. This may indicate a block in the downstream glycerolipid metabolism pathway in GPD3 overexpression lines. Thus, lipid engineering by GPDH modification may depend on the activities of other downstream enzyme steps. These results also suggest that GPD2 and GPD3 GPDH isoforms are important for nutrient starvation-induced TAG accumulation but have distinct metabolic functions.
© 2017 The author(s). All Rights Reserved.
Figures
Similar articles
-
A multidomain enzyme, with glycerol-3-phosphate dehydrogenase and phosphatase activities, is involved in a chloroplastic pathway for glycerol synthesis in Chlamydomonas reinhardtii.Plant J. 2017 Jun;90(6):1079-1092. doi: 10.1111/tpj.13530. Epub 2017 Apr 11. Plant J. 2017. PMID: 28273364
-
In silico cloning and characterization of the glycerol-3-phosphate dehydrogenase (GPDH) gene family in the green microalga Chlamydomonas reinhardtii.Curr Microbiol. 2012 May;64(5):477-85. doi: 10.1007/s00284-012-0095-6. Epub 2012 Feb 23. Curr Microbiol. 2012. PMID: 22358185
-
Anaerobic and aerobic batch cultivations of Saccharomyces cerevisiae mutants impaired in glycerol synthesis.Yeast. 2000 Mar 30;16(5):463-74. doi: 10.1002/(SICI)1097-0061(20000330)16:5<463::AID-YEA535>3.0.CO;2-3. Yeast. 2000. PMID: 10705374
-
The response to inositol: regulation of glycerolipid metabolism and stress response signaling in yeast.Chem Phys Lipids. 2014 May;180:23-43. doi: 10.1016/j.chemphyslip.2013.12.013. Epub 2014 Jan 10. Chem Phys Lipids. 2014. PMID: 24418527 Free PMC article. Review.
-
Implications of glycerol metabolism for lipid production.Prog Lipid Res. 2017 Oct;68:12-25. doi: 10.1016/j.plipres.2017.07.002. Epub 2017 Aug 1. Prog Lipid Res. 2017. PMID: 28778473 Review.
Cited by
-
Potassium channel KCN11 is required for maintaining cellular osmolarity during nitrogen starvation to control proper cell physiology and TAG accumulation in Chlamydomonas reinhardtii.Biotechnol Biofuels. 2020 Jul 20;13:129. doi: 10.1186/s13068-020-01769-x. eCollection 2020. Biotechnol Biofuels. 2020. PMID: 32699552 Free PMC article.
-
Effects of the skp1 gene of the SCF complex on lipid metabolism and response to abiotic stress in Chlamydomonas reinhardtii.Front Plant Sci. 2025 Mar 17;16:1527439. doi: 10.3389/fpls.2025.1527439. eCollection 2025. Front Plant Sci. 2025. PMID: 40166727 Free PMC article.
-
Integrating Transcriptomics and Metabolomics to Characterize Metabolic Regulation to Elevated CO2 in Chlamydomonas Reinhardtii.Mar Biotechnol (NY). 2021 Apr;23(2):255-275. doi: 10.1007/s10126-021-10021-y. Epub 2021 Mar 10. Mar Biotechnol (NY). 2021. PMID: 33689052
-
Jasmonates and Ethylene Shape Floridoside Synthesis during Carposporogenesis in the Red Seaweed Grateloupia imbricata.Mar Drugs. 2024 Feb 28;22(3):115. doi: 10.3390/md22030115. Mar Drugs. 2024. PMID: 38535456 Free PMC article.
-
Novel insights into salinity-induced lipogenesis and carotenogenesis in the oleaginous astaxanthin-producing alga Chromochloris zofingiensis: a multi-omics study.Biotechnol Biofuels. 2020 Apr 16;13:73. doi: 10.1186/s13068-020-01714-y. eCollection 2020. Biotechnol Biofuels. 2020. PMID: 32322303 Free PMC article.
References
-
- André L, Hemming A, Adler L (1991) Osmoregulation in Saccharomyces cerevisiae: studies on the osmotic induction of glycerol production and glycerol-3-phosphate dehydrogenase (NAD+). FEBS Lett 286: 13–17 - PubMed
-
- Bajhaiya AK, Ziehe Moreira J, Pittman JK (2017) Transcriptional engineering of microalgae: prospects for high-value chemicals. Trends Biotechnol 35: 95–99 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

