Linkage mapping of Mungbean yellow mosaic India virus (MYMIV) resistance gene in soybean
- PMID: 28588385
- PMCID: PMC5445968
- DOI: 10.1270/jsbbs.16115
Linkage mapping of Mungbean yellow mosaic India virus (MYMIV) resistance gene in soybean
Abstract
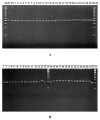
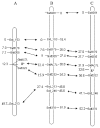
Mungbean Yellow Mosaic India Virus (MYMIV) is one of the most prevalent pathogen that limits soybean production in India. In this study RILs derived from JS335, dominant but MYMIV susceptible variety and PI171443, donor of MYMIV resistance gene in most of the MYMIV resistant varieties released in India and F2 population derived from SL525, a resistant variety released for northern India and NRC101, a susceptible genotype were used to study the inheritance of MYMIV resistance and map the gene responsible for MYMIV resistance. F1s were found to be completely susceptible. F2:3 and RILs population segregated to fit a ratio of 1:2:1 and 1:1 indicating that a single recessive gene controlled resistance to MYMIV. BSA was performed using 144 polymorphic SSR markers. MYMIV resistance gene was mapped on chr 6 (LG C2) within a 3.5-cM genome region between two SSR markers GMAC7L and Satt322 whose size was estimated to be 77.115 kb (position of 12,259,594-12,336,709 bp). This is the first report on linkage mapping of MYMIV resistance gene in soybean. This will be helpful in breeding soybean varieties for resistance against MYMIV responsible for wide spread damage to soybean crop in India using Marker Assisted Selection.
Keywords: MYMIV; SSR; mapping; resistance gene; soybean.
Figures
Similar articles
-
Detection of quantitative trait loci for mungbean yellow mosaic India virus (MYMIV) resistance in mungbean (Vigna radiata (L.) Wilczek) in India and Pakistan.Breed Sci. 2013 Dec;63(4):367-73. doi: 10.1270/jsbbs.63.367. Epub 2013 Dec 1. Breed Sci. 2013. PMID: 24399908 Free PMC article.
-
Gamma-rays and EMS induced resistance to mungbean yellow mosaic India virus in mungbean [Vigna radiata (L.) R. Wilczek] and its validation using linked molecular markers.Int J Radiat Biol. 2022;98(1):69-81. doi: 10.1080/09553002.2022.1998710. Epub 2021 Nov 12. Int J Radiat Biol. 2022. PMID: 34705607
-
A large-effect QTL introgressed from ricebean imparts resistance to Mungbean yellow mosaic India virus in blackgram (Vigna mungo (L.) Hepper).Theor Appl Genet. 2022 Dec;135(12):4495-4506. doi: 10.1007/s00122-022-04234-5. Epub 2022 Oct 22. Theor Appl Genet. 2022. PMID: 36271056
-
Development of infectious clones of mungbean yellow mosaic India virus (MYMIV, Begomovirus vignaradiataindiaense) infecting mungbean [Vigna radiata (L.) R. Wilczek] and evaluation of a RIL population for MYMIV resistance.PLoS One. 2024 Oct 22;19(10):e0310003. doi: 10.1371/journal.pone.0310003. eCollection 2024. PLoS One. 2024. PMID: 39436879 Free PMC article.
-
Yellow Mosaic Disease (YMD) of Mungbean (Vigna radiata (L.) Wilczek): Current Status and Management Opportunities.Front Plant Sci. 2020 Jun 24;11:918. doi: 10.3389/fpls.2020.00918. eCollection 2020. Front Plant Sci. 2020. PMID: 32670329 Free PMC article. Review.
Cited by
-
Identification of Genomic Regions Associated with Vine Growth and Plant Height of Soybean.Int J Mol Sci. 2022 May 22;23(10):5823. doi: 10.3390/ijms23105823. Int J Mol Sci. 2022. PMID: 35628633 Free PMC article.
-
A source of resistance against yellow mosaic disease in soybeans correlates with a novel mutation in a resistance gene.Front Plant Sci. 2023 Nov 24;14:1230559. doi: 10.3389/fpls.2023.1230559. eCollection 2023. Front Plant Sci. 2023. PMID: 38078080 Free PMC article.
-
Expression of short hairpin RNA (shRNA) targeting AC2 gene of Mungbean yellow mosaic India virus (MYMIV) reduces the viral titre in soybean.3 Biotech. 2019 Sep;9(9):334. doi: 10.1007/s13205-019-1865-7. Epub 2019 Aug 16. 3 Biotech. 2019. PMID: 31475086 Free PMC article.
-
Genome-wide association studies for earliness, MYMIV resistance, and other associated traits in mungbean (Vigna radiata L. Wilczek) using genotyping by sequencing approach.PeerJ. 2024 Jan 26;12:e16653. doi: 10.7717/peerj.16653. eCollection 2024. PeerJ. 2024. PMID: 38288464 Free PMC article.
-
Molecular characterization and genetic diversity studies of Indian soybean (Glycine max (L.) Merr.) cultivars using SSR markers.Mol Biol Rep. 2022 Mar;49(3):2129-2140. doi: 10.1007/s11033-021-07030-4. Epub 2021 Dec 11. Mol Biol Rep. 2022. PMID: 34894334 Free PMC article.
References
-
- Doyle, J.J. and Doyle, J.L. (1990) Isolation of plant DNA from fresh tissue. Focus 12: 13–15.
-
- Hanley-Bowdoin, L., Settlage, S.B., Orozco, B.M., Nagar, S. and Robertson, D. (1999) Geminiviruses: models for plant DNA replication, transcription, and cell cycle regulation. Crit. Rev. Biochem. Mol. Biol. 35: 105–140. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
