Long-Term Culture of Self-renewing Pancreatic Progenitors Derived from Human Pluripotent Stem Cells
- PMID: 28591650
- PMCID: PMC5470345
- DOI: 10.1016/j.stemcr.2017.05.019
Long-Term Culture of Self-renewing Pancreatic Progenitors Derived from Human Pluripotent Stem Cells
Abstract
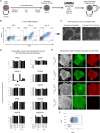
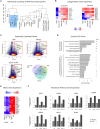
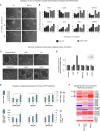
Pluripotent stem cells have been proposed as an unlimited source of pancreatic β cells for studying and treating diabetes. However, the long, multi-step differentiation protocols used to generate functional β cells inevitably exhibit considerable variability, particularly when applied to pluripotent cells from diverse genetic backgrounds. We have developed culture conditions that support long-term self-renewal of human multipotent pancreatic progenitors, which are developmentally more proximal to the specialized cells of the adult pancreas. These cultured pancreatic progenitor (cPP) cells express key pancreatic transcription factors, including PDX1 and SOX9, and exhibit transcriptomes closely related to their in vivo counterparts. Upon exposure to differentiation cues, cPP cells give rise to pancreatic endocrine, acinar, and ductal lineages, indicating multilineage potency. Furthermore, cPP cells generate insulin+ β-like cells in vitro and in vivo, suggesting that they offer a convenient alternative to pluripotent cells as a source of adult cell types for modeling pancreatic development and diabetes.
Keywords: culture conditions; directed differentiation; pancreatic development; pancreatic progenitors; self-renewal; tissue stem cells; β cell differentiation.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Enhanced differentiation of human pluripotent stem cells into pancreatic progenitors co-expressing PDX1 and NKX6.1.Stem Cell Res Ther. 2018 Jan 23;9(1):15. doi: 10.1186/s13287-017-0759-z. Stem Cell Res Ther. 2018. PMID: 29361979 Free PMC article.
-
In Vitro Differentiation of Pluripotent Stem Cells into Functional β Islets Under 2D and 3D Culture Conditions and In Vivo Preclinical Validation of 3D Islets.Methods Mol Biol. 2016;1341:257-84. doi: 10.1007/7651_2015_230. Methods Mol Biol. 2016. PMID: 25783769
-
Small molecule AT7867 proliferates PDX1-expressing pancreatic progenitor cells derived from human pluripotent stem cells.Stem Cell Res. 2017 Oct;24:61-68. doi: 10.1016/j.scr.2017.08.010. Epub 2017 Aug 17. Stem Cell Res. 2017. PMID: 28843156
-
In vitro differentiation and expansion of human pluripotent stem cell-derived pancreatic progenitors.Rev Diabet Stud. 2014 Spring;11(1):19-34. doi: 10.1900/RDS.2014.11.19. Epub 2014 May 10. Rev Diabet Stud. 2014. PMID: 25148365 Free PMC article. Review.
-
In vitro generation of pancreatic β-cells for diabetes treatment. I. β-like cells derived from human pluripotent stem cells.Folia Histochem Cytobiol. 2019;57(1):1-14. doi: 10.5603/FHC.a2019.0001. Epub 2019 Mar 14. Folia Histochem Cytobiol. 2019. PMID: 30869153 Review.
Cited by
-
Reconstructing human pancreatic differentiation by mapping specific cell populations during development.Elife. 2017 Jul 21;6:e27564. doi: 10.7554/eLife.27564. Elife. 2017. PMID: 28731406 Free PMC article.
-
USP7 controls NGN3 stability and pancreatic endocrine lineage development.Nat Commun. 2023 Apr 28;14(1):2457. doi: 10.1038/s41467-023-38146-9. Nat Commun. 2023. PMID: 37117185 Free PMC article.
-
Enhancing human pluripotent stem cell differentiation to cardiomyocytes through cardiac progenitor reseeding and cryopreservation.iScience. 2025 Apr 16;28(5):112452. doi: 10.1016/j.isci.2025.112452. eCollection 2025 May 16. iScience. 2025. PMID: 40454098 Free PMC article.
-
Differentiation of beta-like cells from human induced pluripotent stem cell-derived pancreatic progenitor organoids.STAR Protoc. 2022 Aug 30;3(3):101656. doi: 10.1016/j.xpro.2022.101656. eCollection 2022 Sep 16. STAR Protoc. 2022. PMID: 36092820 Free PMC article.
-
Human expandable pancreatic progenitor-derived β cells ameliorate diabetes.Sci Adv. 2022 Feb 25;8(8):eabk1826. doi: 10.1126/sciadv.abk1826. Epub 2022 Feb 23. Sci Adv. 2022. PMID: 35196077 Free PMC article.
References
-
- Barker N., van Es J.H., Kuipers J., Kujala P., van den Born M., Cozijnsen M., Haegebarth A., Korving J., Begthel H., Peters P.J., Clevers H. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature. 2007;449:1003–1007. - PubMed
-
- Barker N., Huch M., Kujala P., van de Wetering M., Snippert H.J., van Es J.H., Sato T., Stange D.E., Begthel H., van den Born M. LGR5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell. 2010;6:25–36. - PubMed
-
- Bhushan A., Itoh N., Kato S., Thiery J.P., Czernichow P., Bellusci S., Scharfmann R. Fgf10 is essential for maintaining the proliferative capacity of epithelial progenitor cells during early pancreatic organogenesis. Development. 2001;128:5109–5117. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

