mRNA quantification using single-molecule FISH in Drosophila embryos
- PMID: 28594816
- PMCID: PMC6668020
- DOI: 10.1038/nprot.2017.030
mRNA quantification using single-molecule FISH in Drosophila embryos
Abstract
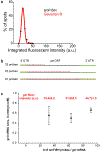
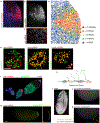
Spatial information is critical to the interrogation of developmental and tissue-level regulation of gene expression. However, this information is usually lost when global mRNA levels from tissues are measured using reverse transcriptase PCR, microarray analysis or high-throughput sequencing. By contrast, single-molecule fluorescence in situ hybridization (smFISH) preserves the spatial information of the cellular mRNA content with subcellular resolution within tissues. Here we describe an smFISH protocol that allows for the quantification of single mRNAs in Drosophila embryos, using commercially available smFISH probes (e.g., short fluorescently labeled DNA oligonucleotides) in combination with wide-field epifluorescence, confocal or instant structured illumination microscopy (iSIM, a super-resolution imaging approach) and a spot-detection algorithm. Fixed Drosophila embryos are hybridized in solution with a mixture of smFISH probes, mounted onto coverslips and imaged in 3D. Individual fluorescently labeled mRNAs are then localized within tissues and counted using spot-detection software to generate quantitative, spatially resolved gene expression data sets. With minimum guidance, a graduate student can successfully implement this protocol. The smFISH procedure described here can be completed in 4-5 d.
Figures



References
-
- Femino AM, Fay FS, Fogarty K & Singer RH Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

