Diacetyl/l-Xylulose Reductase Mediates Chemical Redox Cycling in Lung Epithelial Cells
- PMID: 28595002
- PMCID: PMC5708134
- DOI: 10.1021/acs.chemrestox.7b00052
Diacetyl/l-Xylulose Reductase Mediates Chemical Redox Cycling in Lung Epithelial Cells
Abstract
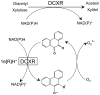
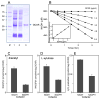
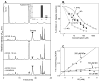
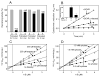
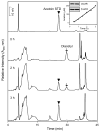
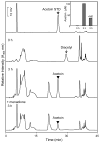
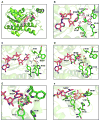
Reactive carbonyls such as diacetyl (2,3-butanedione) and 2,3-pentanedione in tobacco and many food and consumer products are known to cause severe respiratory diseases. Many of these chemicals are detoxified by carbonyl reductases in the lung, in particular, dicarbonyl/l-xylulose reductase (DCXR), a multifunctional enzyme important in glucose metabolism. DCXR is a member of the short-chain dehydrogenase/reductase (SDR) superfamily. Using recombinant human enzyme, we discovered that DCXR mediates redox cycling of a variety of quinones generating superoxide anion, hydrogen peroxide, and, in the presence of transition metals, hydroxyl radicals. Redox cycling activity preferentially utilized NADH as a cosubstrate and was greatest for 9,10-phenanthrenequinone and 1,2-naphthoquinone, followed by 1,4-naphthoquinone and 2-methyl-1,4-naphthoquinone (menadione). Using 9,10-phenanthrenequinone as the substrate, quinone redox cycling was found to inhibit DCXR reduction of l-xylulose and diacetyl. Competitive inhibition of enzyme activity by the quinone was observed with respect to diacetyl (Ki = 190 μM) and l-xylulose (Ki = 940 μM). Abundant DCXR activity was identified in A549 lung epithelial cells when diacetyl was used as a substrate. Quinones inhibited reduction of this dicarbonyl, causing an accumulation of diacetyl in the cells and culture medium and a decrease in acetoin, the reduced product of diacetyl. The identification of DCXR as an enzyme activity mediating chemical redox cycling suggests that it may be important in generating cytotoxic reactive oxygen species in the lung. These activities, together with the inhibition of dicarbonyl/l-xylulose metabolism by redox-active chemicals, as well as consequent deficiencies in pentose metabolism, are likely to contribute to lung injury following exposure to dicarbonyls and quinones.
Conflict of interest statement
The authors declare no competing financial interest
Figures


Similar articles
-
Quinone and nitrofurantoin redox cycling by recombinant cytochrome b5 reductase.Toxicol Appl Pharmacol. 2018 Nov 15;359:102-107. doi: 10.1016/j.taap.2018.09.011. Epub 2018 Sep 15. Toxicol Appl Pharmacol. 2018. PMID: 30222979 Free PMC article.
-
Sepiapterin reductase mediates chemical redox cycling in lung epithelial cells.J Biol Chem. 2013 Jun 28;288(26):19221-37. doi: 10.1074/jbc.M112.402164. Epub 2013 May 2. J Biol Chem. 2013. PMID: 23640889 Free PMC article.
-
Dicarbonyl/l-xylulose reductase (DCXR): The multifunctional pentosuria enzyme.Int J Biochem Cell Biol. 2013 Nov;45(11):2563-7. doi: 10.1016/j.biocel.2013.08.010. Epub 2013 Aug 27. Int J Biochem Cell Biol. 2013. PMID: 23988570 Review.
-
Human DCXR - another 'moonlighting protein' involved in sugar metabolism, carbonyl detoxification, cell adhesion and male fertility?Biol Rev Camb Philos Soc. 2015 Feb;90(1):254-78. doi: 10.1111/brv.12108. Epub 2014 Apr 10. Biol Rev Camb Philos Soc. 2015. PMID: 24720935 Review.
-
(-)-Epigallocatechin-3-gallate, a potential inhibitor to human dicarbonyl/L-xylulose reductase.J Biochem. 2013 Aug;154(2):167-75. doi: 10.1093/jb/mvt039. Epub 2013 May 9. J Biochem. 2013. PMID: 23661708
Cited by
-
Key co-expressed genes correlated with blood serum parameters of pigs fed with different fatty acid profile diets.Front Genet. 2024 Jul 3;15:1394971. doi: 10.3389/fgene.2024.1394971. eCollection 2024. Front Genet. 2024. PMID: 39021677 Free PMC article.
-
Effects of E-Cigarette Flavoring Chemicals on Human Macrophages and Bronchial Epithelial Cells.Int J Environ Res Public Health. 2021 Oct 22;18(21):11107. doi: 10.3390/ijerph182111107. Int J Environ Res Public Health. 2021. PMID: 34769627 Free PMC article.
-
DCXR promotes cell proliferation by promoting the activity of aerobic glycolysis in breast cancer.Mol Med Rep. 2023 Feb;27(2):31. doi: 10.3892/mmr.2022.12918. Epub 2022 Dec 23. Mol Med Rep. 2023. PMID: 36562355 Free PMC article.
-
Identification of Estrogen-Responsive Proteins in Mouse Seminal Vesicles Through Mass Spectrometry-Based Proteomics.Pharmaceuticals (Basel). 2024 Nov 9;17(11):1508. doi: 10.3390/ph17111508. Pharmaceuticals (Basel). 2024. PMID: 39598420 Free PMC article.
-
A comprehensive bioinformatics analysis of pathways and biomarkers shared between type 2 diabetes mellitus and chronic obstructive pulmonary disease.Front Immunol. 2025 Jul 25;16:1536551. doi: 10.3389/fimmu.2025.1536551. eCollection 2025. Front Immunol. 2025. PMID: 40787468 Free PMC article.
References
-
- Ebert B, Kisiela M, Maser E. Human DCXR - another ’moonlighting protein’ involved in sugar metabolism, carbonyl detoxification, cell adhesion and male fertility? Biol Rev Camb Philos Soc. 2015;90:254–278. - PubMed
-
- Nakagawa J, Ishikura S, Asami J, Isaji T, Usami N, Hara A, Sakurai T, Tsuritani K, Oda K, Takahashi M, Yoshimoto M, Otsuka N, Kitamura K. Molecular characterization of mammalian dicarbonyl/l-xylulose reductase and its localization in kidney. J Biol Chem. 2002;277:17883–17891. - PubMed
-
- Asami J, Odani H, Ishii A, Oide K, Sudo T, Nakamura A, Miyata N, Otsuka N, Maeda K, Nakagawa J. Suppression of AGE precursor formation following unilateral ureteral obstruction in mouse kidneys by transgenic expression of alpha-dicarbonyl/l-xylulose reductase. Biosci Biotechnol, Biochem. 2006;70:2899–2905. - PubMed
-
- Touster O, Hutcheson RM, Rice L. The influence of D-glucuronolactone on the excretion of l-xylulose by humans and guinea pigs. Biochim Biophys Acta. 1955;215:677–684. - PubMed
-
- Winegrad AI, Burden CL. l-xylulose metabolism in diabetes mellitus. N Engl J Med. 1966;274:298–305. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

