Genome-Edited T Cell Therapies
- PMID: 28596938
- PMCID: PMC5445182
- DOI: 10.1007/s40778-017-0077-5
Genome-Edited T Cell Therapies
Abstract
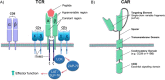
Purpose of review: Alternative approaches to conventional drug-based cancer treatments have seen T cell therapies deployed more widely over the last decade. This is largely due to their ability to target and kill specific cell types based on receptor recognition. Introduction of recombinant T cell receptors (TCRs) using viral vectors and HLA-independent T cell therapies using chimeric antigen receptors (CARs) are discussed. This article reviews the tools used for genome editing, with particular emphasis on the applications of site-specific DNA nuclease mediated editing for T cell therapies.
Recent findings: Genetic engineering of T cells using TCRs and CARs with redirected antigen-targeting specificity has resulted in clinical success of several immunotherapies. In conjunction, the application of genome editing technologies has resulted in the generation of HLA-independent universal T cells for allogeneic transplantation, improved T cell sustainability through knockout of the checkpoint inhibitor, programmed cell death protein-1 (PD-1), and has shown efficacy as an antiviral therapy through direct targeting of viral genomic sequences and entry receptors.
Summary: The combined use of engineered antigen-targeting moieties and innovative genome editing technologies have recently shown success in a small number of clinical trials targeting HIV and hematological malignancies and are now being incorporated into existing strategies for other immunotherapies.
Keywords: CRISPR/Cas9; Chimeric antigen receptors; Clinical trials; Genome editing; Immunotherapy; T cell receptors; T cell therapies.
Conflict of interest statement
Conflict of Interest
Juliette M.K.M. Delhove declares that she has no conflict of interest. Waseem Qasim reports research funding from Cellectis/Servier, Miltenyi, Bellicum, Calimmune, and Autolus and personal fees from Autolus and Orchard. In addition, Dr. Qasim has patents pending for gene-edited T cells.
Human and Animal Rights and Informed Consent
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Figures

Similar articles
-
Genome-edited allogeneic donor "universal" chimeric antigen receptor T cells.Blood. 2023 Feb 23;141(8):835-845. doi: 10.1182/blood.2022016204. Blood. 2023. PMID: 36223560 Free PMC article.
-
Explorations of CRISPR/Cas9 for improving the long-term efficacy of universal CAR-T cells in tumor immunotherapy.Life Sci. 2023 Mar 1;316:121409. doi: 10.1016/j.lfs.2023.121409. Epub 2023 Jan 19. Life Sci. 2023. PMID: 36681183 Review.
-
Gene Therapy with CRISPR/Cas9 Coming to Age for HIV Cure.AIDS Rev. 2017 Oct-Dec;19(3):167-172. AIDS Rev. 2017. PMID: 29019352
-
Genome-Edited T Cell Therapies.Hematol Oncol Clin North Am. 2022 Aug;36(4):729-744. doi: 10.1016/j.hoc.2022.03.006. Epub 2022 Jun 27. Hematol Oncol Clin North Am. 2022. PMID: 35773047 Review.
-
Innovative Strategies of Reprogramming Immune System Cells by Targeting CRISPR/Cas9-Based Genome-Editing Tools: A New Era of Cancer Management.Int J Nanomedicine. 2023 Sep 29;18:5531-5559. doi: 10.2147/IJN.S424872. eCollection 2023. Int J Nanomedicine. 2023. PMID: 37795042 Free PMC article. Review.
Cited by
-
Posttransplant chimeric antigen receptor therapy.Blood. 2018 Mar 8;131(10):1045-1052. doi: 10.1182/blood-2017-08-752121. Epub 2018 Jan 22. Blood. 2018. PMID: 29358181 Free PMC article. Review.
-
Gene editing to enhance the efficacy of cancer cell therapies.Mol Ther. 2021 Nov 3;29(11):3153-3162. doi: 10.1016/j.ymthe.2021.10.001. Epub 2021 Oct 18. Mol Ther. 2021. PMID: 34673274 Free PMC article. Review.
-
Current perspectives on gene therapy and its involvement in curing genetic disorders.Hum Genet. 2025 Jul;144(7):695-713. doi: 10.1007/s00439-025-02757-7. Epub 2025 Jun 18. Hum Genet. 2025. PMID: 40533637 Review.
-
Industrializing engineered autologous T cells as medicines for solid tumours.Nat Rev Drug Discov. 2021 Jun;20(6):476-488. doi: 10.1038/s41573-021-00175-8. Epub 2021 Apr 8. Nat Rev Drug Discov. 2021. PMID: 33833444 Review.
-
CAR-T cell therapy in ovarian cancer: from the bench to the bedside.Oncotarget. 2017 Aug 4;8(38):64607-64621. doi: 10.18632/oncotarget.19929. eCollection 2017 Sep 8. Oncotarget. 2017. PMID: 28969098 Free PMC article. Review.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
