Tespa1 regulates T cell receptor-induced calcium signals by recruiting inositol 1,4,5-trisphosphate receptors
- PMID: 28598420
- PMCID: PMC5472764
- DOI: 10.1038/ncomms15732
Tespa1 regulates T cell receptor-induced calcium signals by recruiting inositol 1,4,5-trisphosphate receptors
Erratum in
-
Author Correction: Tespa1 regulates T cell receptor-induced calcium signals by recruiting inositol 1,4,5-trisphosphate receptors.Nat Commun. 2018 Mar 12;9:16183. doi: 10.1038/ncomms16183. Nat Commun. 2018. PMID: 29528040 Free PMC article.
Abstract
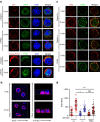
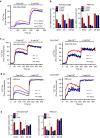
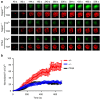
Thymocyte-expressed, positive selection-associated 1 (Tespa1) is important in T cell receptor (TCR)-driven thymocyte development. Downstream of the TCR, Tespa1 is a crucial component of the linker for activation of T cells (LAT) signalosome, facilitating calcium signalling and subsequent MAPK activation. However, it is unknown how Tespa1 elicits calcium signalling. Here, we show that inositol 1,4,5-trisphosphate receptor type 1 (IP3R1) is crucial for Tespa1-optimized, TCR-induced Ca2+ flux and thymocyte development. Upon TCR stimulation, Tespa1 directly interacts with IP3R1 and recruits it to the TCR complex, where IP3R1 is phosphorylated at Y353 by Fyn. This Tespa1-IP3R1 interaction is mediated by the F187 and F188 residues of Tespa1 and the amino-terminus of IP3R1. Tespa1-F187A/F188A mutant mice phenocopy Tespa1-deficient mice with impaired late thymocyte development due to reduced IP3R1 translocation to the TCR-proximal region. Our work elucidates the function of Tespa1 in T cell development and the regulation of TCR-induced Ca2+ signalling through IP3R1.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




References
-
- Malissen B., Gregoire C., Malissen M. & Roncagalli R. Integrative biology of T cell activation. Nat. Immunol. 15, 790–797 (2014). - PubMed
-
- Rao A. & Hogan P. G. Calcium signaling in cells of the immune and hematopoietic systems. Immunol. Rev. 231, 5–9 (2009). - PubMed
-
- Muller M. R. & NFAT Rao A. NFAT, immunity and cancer: a transcription factor comes of age. Nat. Rev. Immunol. 10, 645–656 (2010). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

