Using nanoBRET and CRISPR/Cas9 to monitor proximity to a genome-edited protein in real-time
- PMID: 28600500
- PMCID: PMC5466623
- DOI: 10.1038/s41598-017-03486-2
Using nanoBRET and CRISPR/Cas9 to monitor proximity to a genome-edited protein in real-time
Abstract
Bioluminescence resonance energy transfer (BRET) has been a vital tool for understanding G protein-coupled receptor (GPCR) function. It has been used to investigate GPCR-protein and/or -ligand interactions as well as GPCR oligomerisation. However the utility of BRET is limited by the requirement that the fusion proteins, and in particular the donor, need to be exogenously expressed. To address this, we have used CRISPR/Cas9-mediated homology-directed repair to generate protein-Nanoluciferase (Nluc) fusions under endogenous promotion, thus allowing investigation of proximity between the genome-edited protein and an exogenously expressed protein by BRET. Here we report BRET monitoring of GPCR-mediated β-arrestin2 recruitment and internalisation where the donor luciferase was under endogenous promotion, in live cells and in real time. We have investigated the utility of CRISPR/Cas9 genome editing to create genome-edited fusion proteins that can be used as BRET donors and propose that this strategy can be used to overcome the need for exogenous donor expression.
Conflict of interest statement
C.W.W., H.K.V., H.B.S. and E.K.M.J. have nothing to disclose. K.D.G.P. receives funding from Promega, B.M.G. Labtech and Dimerix as Australian Research Council Linkage Grant participating organisations. These participating organisations played no role in any aspect of the conception or design of the research, collection, analysis and interpretation of results, or writing and editing of the manuscript. K.D.G.P. is Chief Scientific Advisor of Dimerix, of which he maintains a shareholding. Dimerix has proprietary rights to the GPCR-HIT assay.
Figures


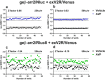


Similar articles
-
NanoBRET ligand binding at a GPCR under endogenous promotion facilitated by CRISPR/Cas9 genome editing.Cell Signal. 2019 Feb;54:27-34. doi: 10.1016/j.cellsig.2018.11.018. Epub 2018 Nov 22. Cell Signal. 2019. PMID: 30471466
-
CRISPR-Mediated Protein Tagging with Nanoluciferase to Investigate Native Chemokine Receptor Function and Conformational Changes.Cell Chem Biol. 2020 May 21;27(5):499-510.e7. doi: 10.1016/j.chembiol.2020.01.010. Epub 2020 Feb 12. Cell Chem Biol. 2020. PMID: 32053779 Free PMC article.
-
NanoBRET: The Bright Future of Proximity-Based Assays.Front Bioeng Biotechnol. 2019 Mar 26;7:56. doi: 10.3389/fbioe.2019.00056. eCollection 2019. Front Bioeng Biotechnol. 2019. PMID: 30972335 Free PMC article. Review.
-
Detection of GPCR/beta-arrestin interactions in live cells using bioluminescence resonance energy transfer technology.Methods Mol Biol. 2009;552:305-17. doi: 10.1007/978-1-60327-317-6_22. Methods Mol Biol. 2009. PMID: 19513659
-
New Horizons on Molecular Pharmacology Applied to Drug Discovery: When Resonance Overcomes Radioligand Binding.Curr Radiopharm. 2017;10(1):16-20. doi: 10.2174/1874471010666170208152420. Curr Radiopharm. 2017. PMID: 28183248 Review.
Cited by
-
Importance of Quantifying Drug-Target Engagement in Cells.ACS Med Chem Lett. 2020 Mar 6;11(4):403-406. doi: 10.1021/acsmedchemlett.9b00570. eCollection 2020 Apr 9. ACS Med Chem Lett. 2020. PMID: 32292539 Free PMC article.
-
Fluorescent ligands: Bringing light to emerging GPCR paradigms.Br J Pharmacol. 2020 Mar;177(5):978-991. doi: 10.1111/bph.14953. Epub 2020 Feb 6. Br J Pharmacol. 2020. PMID: 31877233 Free PMC article. Review.
-
Plasma membrane preassociation drives β-arrestin coupling to receptors and activation.Cell. 2023 May 11;186(10):2238-2255.e20. doi: 10.1016/j.cell.2023.04.018. Epub 2023 May 4. Cell. 2023. PMID: 37146613 Free PMC article.
-
Luminescence- and Fluorescence-Based Complementation Assays to Screen for GPCR Oligomerization: Current State of the Art.Int J Mol Sci. 2019 Jun 17;20(12):2958. doi: 10.3390/ijms20122958. Int J Mol Sci. 2019. PMID: 31213021 Free PMC article. Review.
-
Endogenous Protein Tagging in Human Induced Pluripotent Stem Cells Using CRISPR/Cas9.J Vis Exp. 2018 Aug 25;(138):58130. doi: 10.3791/58130. J Vis Exp. 2018. PMID: 30199041 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

