Computational design of trimeric influenza-neutralizing proteins targeting the hemagglutinin receptor binding site
- PMID: 28604661
- PMCID: PMC5512607
- DOI: 10.1038/nbt.3907
Computational design of trimeric influenza-neutralizing proteins targeting the hemagglutinin receptor binding site
Abstract
Many viral surface glycoproteins and cell surface receptors are homo-oligomers, and thus can potentially be targeted by geometrically matched homo-oligomers that engage all subunits simultaneously to attain high avidity and/or lock subunits together. The adaptive immune system cannot generally employ this strategy since the individual antibody binding sites are not arranged with appropriate geometry to simultaneously engage multiple sites in a single target homo-oligomer. We describe a general strategy for the computational design of homo-oligomeric protein assemblies with binding functionality precisely matched to homo-oligomeric target sites. In the first step, a small protein is designed that binds a single site on the target. In the second step, the designed protein is assembled into a homo-oligomer such that the designed binding sites are aligned with the target sites. We use this approach to design high-avidity trimeric proteins that bind influenza A hemagglutinin (HA) at its conserved receptor binding site. The designed trimers can both capture and detect HA in a paper-based diagnostic format, neutralizes influenza in cell culture, and completely protects mice when given as a single dose 24 h before or after challenge with influenza.
Figures



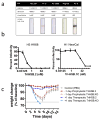
References
-
- Wilson IA, Skehel JJ, Wiley DC. Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 A resolution. Nature. 1981;289:366–373. - PubMed
-
- Heldin CH, Miyazono K, ten Dijke P. TGF-beta signalling from cell membrane to nucleus through SMAD proteins. Nature. 1997;390:465–471. - PubMed
-
- Heldin CH. Dimerization of cell surface receptors in signal transduction. Cell. 1995;80:213–223. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

