Quantifying Abdominal Pigmentation in Drosophila melanogaster
- PMID: 28605370
- PMCID: PMC5608185
- DOI: 10.3791/55732
Quantifying Abdominal Pigmentation in Drosophila melanogaster
Abstract
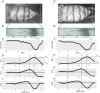
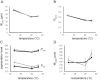
Pigmentation is a morphologically simple but highly variable trait that often has adaptive significance. It has served extensively as a model for understanding the development and evolution of morphological phenotypes. Abdominal pigmentation in Drosophila melanogaster has been particularly useful, allowing researchers to identify the loci that underlie inter- and intraspecific variations in morphology. Hitherto, however, D. melanogaster abdominal pigmentation has been largely assayed qualitatively, through scoring, rather than quantitatively, which limits the forms of statistical analysis that can be applied to pigmentation data. This work describes a new methodology that allows for the quantification of various aspects of the abdominal pigmentation pattern of adult D. melanogaster. The protocol includes specimen mounting, image capture, data extraction, and analysis. All the software used for image capture and analysis feature macros written for open-source image analysis. The advantage of this approach is the ability to precisely measure pigmentation traits using a methodology that is highly reproducible across different imaging systems. While the technique has been used to measure variation in the tergal pigmentation patterns of adult D. melanogaster, the methodology is flexible and broadly applicable to pigmentation patterns in myriad different organisms.
Similar articles
-
Genetic Architecture of Abdominal Pigmentation in Drosophila melanogaster.PLoS Genet. 2015 May 1;11(5):e1005163. doi: 10.1371/journal.pgen.1005163. eCollection 2015 May. PLoS Genet. 2015. PMID: 25933381 Free PMC article.
-
Complex patterns of cis-regulatory polymorphisms in ebony underlie standing pigmentation variation in Drosophila melanogaster.Mol Ecol. 2015 Dec;24(23):5829-41. doi: 10.1111/mec.13432. Epub 2015 Nov 20. Mol Ecol. 2015. PMID: 26503353
-
Phenotypic Plasticity through Transcriptional Regulation of the Evolutionary Hotspot Gene tan in Drosophila melanogaster.PLoS Genet. 2016 Aug 10;12(8):e1006218. doi: 10.1371/journal.pgen.1006218. eCollection 2016 Aug. PLoS Genet. 2016. PMID: 27508387 Free PMC article.
-
Genetic basis of natural variation in body pigmentation in Drosophila melanogaster.Fly (Austin). 2015;9(2):75-81. doi: 10.1080/19336934.2015.1102807. Fly (Austin). 2015. PMID: 26554300 Free PMC article.
-
Bioimage Informatics in the context of Drosophila research.Methods. 2014 Jun 15;68(1):60-73. doi: 10.1016/j.ymeth.2014.04.004. Epub 2014 Apr 13. Methods. 2014. PMID: 24732429 Review.
Cited by
-
Many ways to make darker flies: Intra- and interspecific variation in Drosophila body pigmentation components.Ecol Evol. 2021 May 25;11(12):8136-8155. doi: 10.1002/ece3.7646. eCollection 2021 Jun. Ecol Evol. 2021. PMID: 34188876 Free PMC article.
-
Quantifying Abdominal Coloration of Worker Honey Bees.Insects. 2024 Mar 22;15(4):213. doi: 10.3390/insects15040213. Insects. 2024. PMID: 38667343 Free PMC article.
References
-
- Wittkopp PJ, Beldade P. Development and evolution of insect pigmentation: Genetic mechanisms and the potential consequences of pleiotropy. Semin. Cell Dev. Biol. 2009;20(1):65–71. - PubMed
-
- Monteiro A. Origin, development, and evolution of butterfly eyespots. Annu Rev Entomol. 2015;60:253–271. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases