Flipping states: a few key residues decide the winning conformation of the only universally conserved transcription factor
- PMID: 28605514
- PMCID: PMC5587751
- DOI: 10.1093/nar/gkx523
Flipping states: a few key residues decide the winning conformation of the only universally conserved transcription factor
Abstract
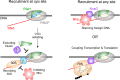
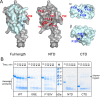
Transcription factors from the NusG family bind to the elongating RNA polymerase to enable synthesis of long RNAs in all domains of life. In bacteria, NusG frequently co-exists with specialized paralogs that regulate expression of a small set of targets, many of which encode virulence factors. Escherichia coli RfaH is the exemplar of this regulatory mechanism. In contrast to NusG, which freely binds to RNA polymerase, RfaH exists in a structurally distinct autoinhibitory state in which the RNA polymerase-binding site is buried at the interface between two RfaH domains. Binding to an ops DNA sequence triggers structural transformation wherein the domains dissociate and RfaH refolds into a NusG-like structure. Formation of the autoinhibitory state, and thus sequence-specific recruitment, represents the decisive step in the evolutionary history of the RfaH subfamily. We used computational and experimental approaches to identify the residues that confer the unique regulatory properties of RfaH. Our analysis highlighted highly conserved Ile and Phe residues at the RfaH interdomain interface. Replacement of these residues with equally conserved Glu and Val counterpart residues in NusG destabilized interactions between the RfaH domains and allowed sequence-independent recruitment to RNA polymerase, suggesting a plausible pathway for diversification of NusG paralogs.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Structural Basis for Transcript Elongation Control by NusG Family Universal Regulators.Cell. 2018 Jun 14;173(7):1650-1662.e14. doi: 10.1016/j.cell.2018.05.017. Epub 2018 Jun 7. Cell. 2018. PMID: 29887376 Free PMC article.
-
Origins and Molecular Evolution of the NusG Paralog RfaH.mBio. 2020 Oct 27;11(5):e02717-20. doi: 10.1128/mBio.02717-20. mBio. 2020. PMID: 33109766 Free PMC article.
-
The universally-conserved transcription factor RfaH is recruited to a hairpin structure of the non-template DNA strand.Elife. 2018 May 9;7:e36349. doi: 10.7554/eLife.36349. Elife. 2018. PMID: 29741479 Free PMC article.
-
NusG-Spt5 Transcription Factors: Universal, Dynamic Modulators of Gene Expression.J Mol Biol. 2025 Jan 1;437(1):168814. doi: 10.1016/j.jmb.2024.168814. Epub 2024 Oct 5. J Mol Biol. 2025. PMID: 39374889 Free PMC article. Review.
-
NusG-Spt5 proteins-Universal tools for transcription modification and communication.Chem Rev. 2013 Nov 13;113(11):8604-19. doi: 10.1021/cr400064k. Epub 2013 May 2. Chem Rev. 2013. PMID: 23638618 Free PMC article. Review. No abstract available.
Cited by
-
NusG, an Ancient Yet Rapidly Evolving Transcription Factor.Front Microbiol. 2021 Jan 8;11:619618. doi: 10.3389/fmicb.2020.619618. eCollection 2020. Front Microbiol. 2021. PMID: 33488562 Free PMC article. Review.
-
Ancient Transcription Factors in the News.mBio. 2019 Feb 26;10(1):e01547-18. doi: 10.1128/mBio.01547-18. mBio. 2019. PMID: 30808693 Free PMC article. Review.
-
Fluid protein fold space and its implications.Bioessays. 2023 Sep;45(9):e2300057. doi: 10.1002/bies.202300057. Epub 2023 Jul 11. Bioessays. 2023. PMID: 37431685 Free PMC article.
-
Metamorphic proteins and how to find them.Curr Opin Struct Biol. 2024 Jun;86:102807. doi: 10.1016/j.sbi.2024.102807. Epub 2024 Mar 26. Curr Opin Struct Biol. 2024. PMID: 38537533 Free PMC article. Review.
-
Identification of Four Amoebicidal Nontoxic Compounds by a Molecular Docking Screen of Naegleria fowleri Sterol Δ8-Δ7-Isomerase and Phenotypic Assays.ACS Infect Dis. 2019 Dec 13;5(12):2029-2038. doi: 10.1021/acsinfecdis.9b00227. Epub 2019 Oct 17. ACS Infect Dis. 2019. PMID: 31583882 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

