The SaeRS Two-Component System Controls Survival of Staphylococcus aureus in Human Blood through Regulation of Coagulase
- PMID: 28611950
- PMCID: PMC5447086
- DOI: 10.3389/fcimb.2017.00204
The SaeRS Two-Component System Controls Survival of Staphylococcus aureus in Human Blood through Regulation of Coagulase
Abstract
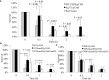
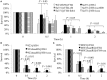
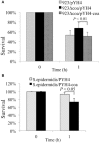
The SaeRS two-component system plays important roles in regulation of key virulence factors and pathogenicity. In this study, however, we found that the deletion mutation of saeRS enhanced bacterial survival in human blood, whereas complementation of the mutant with SaeRS returned survival to wild-type levels. Moreover, these phenomena were observed in different MRSA genetic background isolates, including HA-MRSA WCUH29, CA-MRSA 923, and MW2. To elucidate which gene(s) regulated by SaeRS contribute to the effect, we conducted a series of complementation studies with selected known SaeRS target genes in trans. We found coagulase complementation abolished the enhanced survival of the SaeRS mutant in human blood. The coa and saeRS deletion mutants exhibited a similar survival phenotype in blood. Intriguingly, heterologous expression of coagulase decreased survival of S. epidermidis in human blood. Further, the addition of recombinant coagulase to blood significantly decreased the survival of S. aureus. Further, analysis revealed staphylococcal resistance to killing by hydrogen peroxide was partially dependent on the presence or absence of coagulase. Furthermore, complementation with coagulase, but not SaeRS, returned saeRS/coa double mutant survival in blood to wild-type levels. These data indicate SaeRS modulates bacterial survival in blood in coagulase-dependent manner. Our results provide new insights into the role of staphylococcal SaeRS and coagulase on bacterial survival in human blood.
Keywords: S. aureus; SaeRS; coagulase; survival; two-component system.
Figures



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

