Generation of a microglial developmental index in mice and in humans reveals a sex difference in maturation and immune reactivity
- PMID: 28618077
- PMCID: PMC5540146
- DOI: 10.1002/glia.23176
Generation of a microglial developmental index in mice and in humans reveals a sex difference in maturation and immune reactivity
Erratum in
-
Generation of a microglial developmental index in mice and in humans reveals a sex difference in maturation and immune reactivity.Glia. 2018 Feb;66(2):460. doi: 10.1002/glia.23277. Glia. 2018. PMID: 29230901 No abstract available.
Abstract
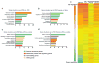
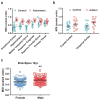
Evidence suggests many neurological disorders emerge when normal neurodevelopmental trajectories are disrupted, i.e., when circuits or cells do not reach their fully mature state. Microglia play a critical role in normal neurodevelopment and are hypothesized to contribute to brain disease. We used whole transcriptome profiling with Next Generation sequencing of purified developing microglia to identify a microglial developmental gene expression program involving thousands of genes whose expression levels change monotonically (up or down) across development. Importantly, the gene expression program was delayed in males relative to females and exposure of adult male mice to LPS, a potent immune activator, accelerated microglial development in males. Next, a microglial developmental index (MDI) generated from gene expression patterns obtained from purified mouse microglia, was applied to human brain transcriptome datasets to test the hypothesis that variability in microglial development is associated with human diseases such as Alzheimer's and autism where microglia have been suggested to play a role. MDI was significantly increased in both Alzheimer's Disease and in autism, suggesting that accelerated microglial development may contribute to neuropathology. In conclusion, we identified a microglia-specific gene expression program in mice that was used to create a microglia developmental index, which was applied to human datasets containing heterogeneous cell types to reveal differences between healthy and diseased brain samples, and between males and females. This powerful tool has wide ranging applicability to examine microglial development within the context of disease and in response to other variables such as stress and pharmacological treatments.
Keywords: development; microglia; sex differences; whole transcriptome analysis.
© 2017 Wiley Periodicals, Inc.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




References
-
- Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, … Barres BA. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci. 2008;28(1):264–278. doi: 10.1523/JNEUROSCI.4178-07.2008. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

