Structural Basis of Egg Coat-Sperm Recognition at Fertilization
- PMID: 28622512
- PMCID: PMC5480393
- DOI: 10.1016/j.cell.2017.05.033
Structural Basis of Egg Coat-Sperm Recognition at Fertilization
Abstract
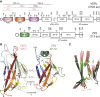
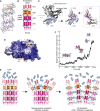
Recognition between sperm and the egg surface marks the beginning of life in all sexually reproducing organisms. This fundamental biological event depends on the species-specific interaction between rapidly evolving counterpart molecules on the gametes. We report biochemical, crystallographic, and mutational studies of domain repeats 1-3 of invertebrate egg coat protein VERL and their interaction with cognate sperm protein lysin. VERL repeats fold like the functionally essential N-terminal repeat of mammalian sperm receptor ZP2, whose structure is also described here. Whereas sequence-divergent repeat 1 does not bind lysin, repeat 3 binds it non-species specifically via a high-affinity, largely hydrophobic interface. Due to its intermediate binding affinity, repeat 2 selectively interacts with lysin from the same species. Exposure of a highly positively charged surface of VERL-bound lysin suggests that complex formation both disrupts the organization of egg coat filaments and triggers their electrostatic repulsion, thereby opening a hole for sperm penetration and fusion.
Keywords: X-ray crystallography; biological evolution; fertilization; protein interaction domains and motifs; sperm-ovum interactions; zona pellucida; zona pellucida glycoproteins.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures













Comment in
-
Sex at Atomic Resolution.Cell. 2017 Jun 15;169(7):1174-1176. doi: 10.1016/j.cell.2017.05.043. Cell. 2017. PMID: 28622504
Similar articles
-
ZP domain proteins in the abalone egg coat include a paralog of VERL under positive selection that binds lysin and 18-kDa sperm proteins.Mol Biol Evol. 2010 Jan;27(1):193-203. doi: 10.1093/molbev/msp221. Mol Biol Evol. 2010. PMID: 19767347 Free PMC article.
-
Crystal structure of the ZP-N domain of ZP3 reveals the core fold of animal egg coats.Nature. 2008 Dec 4;456(7222):653-7. doi: 10.1038/nature07599. Nature. 2008. PMID: 19052627
-
Egg Coat Proteins Across Metazoan Evolution.Curr Top Dev Biol. 2018;130:443-488. doi: 10.1016/bs.ctdb.2018.03.005. Epub 2018 May 7. Curr Top Dev Biol. 2018. PMID: 29853187 Free PMC article. Review.
-
Structure of Zona Pellucida Module Proteins.Curr Top Dev Biol. 2018;130:413-442. doi: 10.1016/bs.ctdb.2018.02.007. Epub 2018 May 10. Curr Top Dev Biol. 2018. PMID: 29853186 Review.
-
Concerted evolution in an egg receptor for a rapidly evolving abalone sperm protein.Science. 1998 Jul 31;281(5377):710-2. doi: 10.1126/science.281.5377.710. Science. 1998. PMID: 9685267
Cited by
-
Female fertility and the mammalian egg's zona pellucida.Histol Histopathol. 2024 Oct;39(10):1273-1284. doi: 10.14670/HH-18-728. Epub 2024 Mar 4. Histol Histopathol. 2024. PMID: 38487866 Review.
-
The conserved fertility factor SPACA4/Bouncer has divergent modes of action in vertebrate fertilization.Proc Natl Acad Sci U S A. 2021 Sep 28;118(39):e2108777118. doi: 10.1073/pnas.2108777118. Proc Natl Acad Sci U S A. 2021. PMID: 34556579 Free PMC article.
-
Structural basis of ZP2-targeted female nonhormonal contraception.Proc Natl Acad Sci U S A. 2025 Apr 15;122(15):e2426057122. doi: 10.1073/pnas.2426057122. Epub 2025 Apr 11. Proc Natl Acad Sci U S A. 2025. PMID: 40215272 Free PMC article.
-
Recurrent Co-Option and Recombination of Cytokine and Three Finger Proteins in Multiple Reproductive Tissues Throughout Salamander Evolution.Front Cell Dev Biol. 2022 Feb 23;10:828947. doi: 10.3389/fcell.2022.828947. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35281090 Free PMC article.
-
Discovery of archaeal fusexins homologous to eukaryotic HAP2/GCS1 gamete fusion proteins.Nat Commun. 2022 Jul 6;13(1):3880. doi: 10.1038/s41467-022-31564-1. Nat Commun. 2022. PMID: 35794124 Free PMC article.
References
-
- Abrahams J.P., Leslie A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 1996;52:30–42. - PubMed
-
- Agirre J., Iglesias-Fernández J., Rovira C., Davies G.J., Wilson K.S., Cowtan K.D. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 2015;22:833–834. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

