Natural DNA Transformation Is Functional in Lactococcus lactis subsp. cremoris KW2
- PMID: 28625996
- PMCID: PMC5541205
- DOI: 10.1128/AEM.01074-17
Natural DNA Transformation Is Functional in Lactococcus lactis subsp. cremoris KW2
Abstract
Lactococcus lactis is one of the most commonly used lactic acid bacteria in the dairy industry. Activation of competence for natural DNA transformation in this species would greatly improve the selection of novel strains with desired genetic traits. Here, we investigated the activation of natural transformation in L. lactis subsp. cremoris KW2, a strain of plant origin whose genome encodes the master competence regulator ComX and the complete set of proteins usually required for natural transformation. In the absence of knowledge about competence regulation in this species, we constitutively overproduced ComX in a reporter strain of late competence phase activation and showed, by transcriptomic analyses, a ComX-dependent induction of all key competence genes. We further demonstrated that natural DNA transformation is functional in this strain and requires the competence DNA uptake machinery. Since constitutive ComX overproduction is unstable, we alternatively expressed comX under the control of an endogenous xylose-inducible promoter. This regulated system was used to successfully inactivate the adaptor protein MecA and subunits of the Clp proteolytic complex, which were previously shown to be involved in ComX degradation in streptococci. In the presence of a small amount of ComX, the deletion of mecA, clpC, or clpP genes markedly increased the activation of the late competence phase and transformability. Altogether, our results report the functionality of natural DNA transformation in L. lactis and pave the way for the identification of signaling mechanisms that trigger the competence state in this species.IMPORTANCE Lactococcus lactis is a lactic acid bacterium of major importance, which is used as a starter species for milk fermentation, a host for heterologous protein production, and a delivery platform for therapeutic molecules. Here, we report the functionality of natural transformation in L. lactis subsp. cremoris KW2 by the overproduction of the master competence regulator ComX. The developed procedure enables a flexible approach to modify the chromosome with single point mutation, sequence insertion, or sequence replacement. These results represent an important step for the genetic engineering of L. lactis that will facilitate the design of strains optimized for industrial applications. This will also help to discover natural regulatory mechanisms controlling competence in the genus Lactococcus.
Keywords: Clp protease; ComX; Lactococcus; competence; lactic acid bacteria; natural DNA transformation; sigma factor.
Copyright © 2017 American Society for Microbiology.
Figures

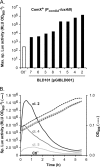
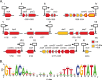



References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

