Cardiac fibroblast transcriptome analyses support a role for interferogenic, profibrotic, and inflammatory genes in anti-SSA/Ro-associated congenital heart block
- PMID: 28626076
- PMCID: PMC5625175
- DOI: 10.1152/ajpheart.00256.2017
Cardiac fibroblast transcriptome analyses support a role for interferogenic, profibrotic, and inflammatory genes in anti-SSA/Ro-associated congenital heart block
Abstract
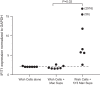
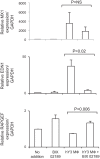
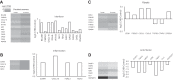
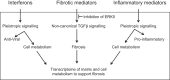
The signature lesion of SSA/Ro autoantibody-associated congenital heart block (CHB) is fibrosis and a macrophage infiltrate, supporting an experimental focus on cues influencing the fibroblast component. The transcriptomes of human fetal cardiac fibroblasts were analyzed using two complementary approaches. Cardiac injury conditions were simulated in vitro by incubating human fetal cardiac fibroblasts with supernatants from macrophages transfected with the SSA/Ro-associated noncoding Y ssRNA. The top 10 upregulated transcripts in the stimulated fibroblasts reflected a type I interferon (IFN) response [e.g., IFN-induced protein 44-like (IFI44L), of MX dynamin-like GTPase (MX)1, MX2, and radical S-adenosyl methionine domain containing 2 (Rsad2)]. Within the fibrotic pathway, transcript levels of endothelin-1 (EDN1), phosphodiesterase (PDE)4D, chemokine (C-X-C motif) ligand (CXCL)2, and CXCL3 were upregulated, while others, including adenomedullin, RAP guanine nucleotide exchange factor 3 (RAPGEF3), tissue inhibitor of metalloproteinase (TIMP)1, TIMP3, and dual specificity phosphatase 1, were downregulated. Agnostic Database for Annotation, Visualization and Integrated Discovery analysis revealed a significant increase in inflammatory genes, including complement C3A receptor 1 (C3AR1), F2R-like thrombin/trypsin receptor 3, and neutrophil cytosolic factor 2. In addition, stimulated fibroblasts expressed high levels of phospho-MADS box transcription enhancer factor 2 [a substrate of MAPK5 (ERK5)], which was inhibited by BIX-02189, a specific inhibitor of ERK5. Translation to human disease leveraged an unprecedented opportunity to interrogate the transcriptome of fibroblasts freshly isolated and cell sorted without stimulation from a fetal heart with CHB and a matched healthy heart. Consistent with the in vitro data, five IFN response genes were among the top 10 most highly expressed transcripts in CHB fibroblasts. In addition, the expression of matrix-related genes reflected fibrosis. These data support the novel finding that cardiac injury in CHB may occur secondary to abnormal remodeling due in part to upregulation of type 1 IFN response genes.NEW & NOTEWORTHY Congenital heart block is a rare disease of the fetal heart associated with maternal anti-Ro autoantibodies which can result in death and for survivors, lifelong pacing. This study provides in vivo and in vitro transcriptome-support that injury may be mediated by an effect of Type I Interferon on fetal fibroblasts.
Keywords: RNASeq; autoimmune congenital heart block; cardiac fibroblast.
Copyright © 2017 the American Physiological Society.
Figures



Similar articles
-
Siglec-1 Macrophages and the Contribution of IFN to the Development of Autoimmune Congenital Heart Block.J Immunol. 2019 Jan 1;202(1):48-55. doi: 10.4049/jimmunol.1800357. Epub 2018 Dec 5. J Immunol. 2019. PMID: 30518570 Free PMC article.
-
Ro60-associated single-stranded RNA links inflammation with fetal cardiac fibrosis via ligation of TLRs: a novel pathway to autoimmune-associated heart block.J Immunol. 2010 Feb 15;184(4):2148-55. doi: 10.4049/jimmunol.0902248. Epub 2010 Jan 20. J Immunol. 2010. PMID: 20089705 Free PMC article.
-
Role of hypoxia and cAMP in the transdifferentiation of human fetal cardiac fibroblasts: implications for progression to scarring in autoimmune-associated congenital heart block.Arthritis Rheum. 2007 Dec;56(12):4120-31. doi: 10.1002/art.23061. Arthritis Rheum. 2007. PMID: 18050204
-
Dying right to live longer: positing apoptosis as a link between maternal autoantibodies and congenital heart block.Lupus. 2008 Feb;17(2):86-90. doi: 10.1177/0961203307085115. Lupus. 2008. PMID: 18250129 Review.
-
Autoimmune-associated congenital heart block: dissecting the cascade from immunologic insult to relentless fibrosis.Anat Rec A Discov Mol Cell Evol Biol. 2004 Oct;280(2):1027-35. doi: 10.1002/ar.a.20072. Anat Rec A Discov Mol Cell Evol Biol. 2004. PMID: 15368347 Review.
Cited by
-
Siglec-1 Macrophages and the Contribution of IFN to the Development of Autoimmune Congenital Heart Block.J Immunol. 2019 Jan 1;202(1):48-55. doi: 10.4049/jimmunol.1800357. Epub 2018 Dec 5. J Immunol. 2019. PMID: 30518570 Free PMC article.
-
Molecular Mechanisms of Fetal and Neonatal Lupus: A Narrative Review of an Autoimmune Disease Transferal across the Placenta.Int J Mol Sci. 2024 May 10;25(10):5224. doi: 10.3390/ijms25105224. Int J Mol Sci. 2024. PMID: 38791261 Free PMC article. Review.
-
Maternal Immune Activation: Implications for Congenital Heart Defects.Clin Rev Allergy Immunol. 2025 Apr 2;68(1):36. doi: 10.1007/s12016-025-09049-y. Clin Rev Allergy Immunol. 2025. PMID: 40175706 Review.
-
Cell atlas of the foetal human heart and implications for autoimmune-mediated congenital heart block.Cardiovasc Res. 2020 Jul 1;116(8):1446-1457. doi: 10.1093/cvr/cvz257. Cardiovasc Res. 2020. PMID: 31589297 Free PMC article.
-
Common innate pathways to autoimmune disease.Clin Immunol. 2020 Mar;212:108361. doi: 10.1016/j.clim.2020.108361. Epub 2020 Feb 10. Clin Immunol. 2020. PMID: 32058071 Free PMC article. Review.
References
-
- Alric L, Duffaut M, Selves J, Sandre K, Mularczyck M, Izopet J, Desmorat H, Bureau C, Chaouche N, Dalbergue B, Vinel JP. Maintenance therapy with gradual reduction of the interferon dose over one year improves histological response in patients with chronic hepatitis C with biochemical response: results of a randomized trial. J Hepatol 35: 272–278, 2001. doi:10.1016/S0168-8278(01)00110-6. - DOI - PubMed
-
- Alvarez D, Briassouli P, Clancy RM, Zavadil J, Reed JH, Abellar RG, Halushka M, Fox-Talbot K, Barrat FJ, Buyon JP. A novel role of endothelin-1 in linking Toll-like receptor 7-mediated inflammation to fibrosis in congenital heart block. J Biol Chem 286: 30444–30454, 2011. doi:10.1074/jbc.M111.263657. - DOI - PMC - PubMed
-
- Azuma A, Li YJ, Abe S, Usuki J, Matsuda K, Henmi S, Miyauchi Y, Ueda K, Izawa A, Sone S, Hashimoto S, Kudoh S. Interferon-β inhibits bleomycin-induced lung fibrosis by decreasing transforming growth factor-β and thrombospondin. Am J Respir Cell Mol Biol 32: 93–98, 2005. doi:10.1165/rcmb.2003-0374OC. - DOI - PubMed
-
- Bhandare R, Damera G, Banerjee A, Flammer JR, Keslacy S, Rogatsky I, Panettieri RA, Amrani Y, Tliba O. Glucocorticoid receptor interacting protein-1 restores glucocorticoid responsiveness in steroid-resistant airway structural cells. Am J Respir Cell Mol Biol 42: 9–15, 2010. doi:10.1165/rcmb.2009-0239RC. - DOI - PMC - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

