Assembly mechanism of the α-pore-forming toxin cytolysin A from Escherichia coli
- PMID: 28630151
- PMCID: PMC5483514
- DOI: 10.1098/rstb.2016.0211
Assembly mechanism of the α-pore-forming toxin cytolysin A from Escherichia coli
Abstract
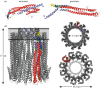
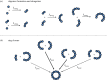
The cytolytic toxin cytolysin A (ClyA) from Escherichia coli is probably one of the best-characterized examples of bacterial, α-pore-forming toxins (α-PFTs). Like other PFTs, ClyA exists in a soluble, monomeric form that assembles to an annular, homo-oligomeric pore complex upon contact with detergent or target membranes. Comparison of the three-dimensional structures of the 34 kDa monomer and the protomer in the context of the dodecameric pore complex revealed that ClyA undergoes one of the largest conformational transitions described for proteins so far, in which 55% of the residues change their position and 16% of the residues adopt a different secondary structure in the protomer. Studies on the assembly of ClyA revealed a unique mechanism that differs from the assembly mechanism of other PFTs. The rate-liming step of pore formation proved to be the unimolecular conversion of the monomer to an assembly-competent protomer, during which a molten globule-like off-pathway intermediate accumulates. The oligomerization of protomers to pore complexes is fast and follows a kinetic scheme in which mixtures of linear oligomers of different size are formed first, followed by very rapid and specific association of pairs of oligomers that can directly perform ring closure to the dodecameric pore complex.This article is part of the themed issue 'Membrane pores: from structure and assembly, to medicine and technology'.
Keywords: assembly kinetics; assembly of membrane complexes; cytolysin A; α-pore-forming toxins.
© 2017 The Author(s).
Conflict of interest statement
We declare we have no competing interests.
Figures

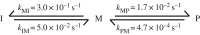
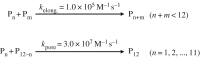


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

