A liver-specific gene expression panel predicts the differentiation status of in vitro hepatocyte models
- PMID: 28640507
- PMCID: PMC5698781
- DOI: 10.1002/hep.29324
A liver-specific gene expression panel predicts the differentiation status of in vitro hepatocyte models
Abstract
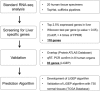
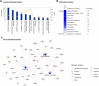
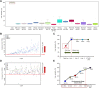
Alternative cell sources, such as three-dimensional organoids and induced pluripotent stem cell-derived cells, might provide a potentially effective approach for both drug development applications and clinical transplantation. For example, the development of cell sources for liver cell-based therapy has been increasingly needed, and liver transplantation is performed for the treatment for patients with severe end-stage liver disease. Differentiated liver cells and three-dimensional organoids are expected to provide new cell sources for tissue models and revolutionary clinical therapies. However, conventional experimental methods confirming the expression levels of liver-specific lineage markers cannot provide complete information regarding the differentiation status or degree of similarity between liver and differentiated cell sources. Therefore, in this study, to overcome several issues associated with the assessment of differentiated liver cells and organoids, we developed a liver-specific gene expression panel (LiGEP) algorithm that presents the degree of liver similarity as a "percentage." We demonstrated that the percentage calculated using the LiGEP algorithm was correlated with the developmental stages of in vivo liver tissues in mice, suggesting that LiGEP can correctly predict developmental stages. Moreover, three-dimensional cultured HepaRG cells and human pluripotent stem cell-derived hepatocyte-like cells showed liver similarity scores of 59.14% and 32%, respectively, although general liver-specific markers were detected.
Conclusion: Our study describes a quantitative and predictive model for differentiated samples, particularly liver-specific cells or organoids; and this model can be further expanded to various tissue-specific organoids; our LiGEP can provide useful information and insights regarding the differentiation status of in vitro liver models. (Hepatology 2017;66:1662-1674).
© 2017 by the American Association for the Study of Liver Diseases.
Figures



Similar articles
-
Feeder-free and serum-free production of hepatocytes, cholangiocytes, and their proliferating progenitors from human pluripotent stem cells: application to liver-specific functional and cytotoxic assays.Cell Reprogram. 2012 Apr;14(2):171-85. doi: 10.1089/cell.2011.0064. Epub 2012 Mar 2. Cell Reprogram. 2012. PMID: 22384928
-
Hepatic differentiation of human pluripotent stem cells on human liver progenitor HepaRG-derived acellular matrix.Exp Cell Res. 2016 Feb 15;341(2):207-17. doi: 10.1016/j.yexcr.2016.02.006. Epub 2016 Feb 8. Exp Cell Res. 2016. PMID: 26854693
-
Transdifferentiation of hepatocyte-like cells from the human hepatoma HepaRG cell line through bipotent progenitor.Hepatology. 2007 Apr;45(4):957-67. doi: 10.1002/hep.21536. Hepatology. 2007. PMID: 17393521
-
Advanced Techniques and Awaited Clinical Applications for Human Pluripotent Stem Cell Differentiation into Hepatocytes.Hepatology. 2021 Aug;74(2):1101-1116. doi: 10.1002/hep.31705. Epub 2021 Aug 22. Hepatology. 2021. PMID: 33420753 Free PMC article. Review.
-
[Robust differentiation of fetal hepatocytes from human embryonic stem cells and iPS].Med Sci (Paris). 2010 Dec;26(12):1061-6. doi: 10.1051/medsci/201026121061. Med Sci (Paris). 2010. PMID: 21187045 Review. French.
Cited by
-
Pluripotent Stem Cell-Derived Hepatocyte-like Cells: Induction Methods and Applications.Int J Mol Sci. 2023 Jul 18;24(14):11592. doi: 10.3390/ijms241411592. Int J Mol Sci. 2023. PMID: 37511351 Free PMC article. Review.
-
Host cell-dependent late entry step as determinant of hepatitis B virus infection.PLoS Pathog. 2022 Jun 17;18(6):e1010633. doi: 10.1371/journal.ppat.1010633. eCollection 2022 Jun. PLoS Pathog. 2022. PMID: 35714170 Free PMC article.
-
Integrative analysis of single-cell RNA-seq and ATAC-seq reveals heterogeneity of induced pluripotent stem cell-derived hepatic organoids.iScience. 2023 Aug 18;26(9):107675. doi: 10.1016/j.isci.2023.107675. eCollection 2023 Sep 15. iScience. 2023. PMID: 37680467 Free PMC article.
-
Transcriptome Classification Reveals Molecular Subgroups in Patients with Hepatitis B Virus.Comput Math Methods Med. 2021 Mar 30;2021:5543747. doi: 10.1155/2021/5543747. eCollection 2021. Comput Math Methods Med. 2021. PMID: 33859718 Free PMC article.
-
Integrating bulk and single-cell RNA sequencing reveals cellular heterogeneity and immune infiltration in hepatocellular carcinoma.Mol Oncol. 2022 Jun;16(11):2195-2213. doi: 10.1002/1878-0261.13190. Epub 2022 Mar 1. Mol Oncol. 2022. PMID: 35124891 Free PMC article.
References
-
- Fennema E, Rivron N, Rouwkema J, van Blitterswijk C, de Boer J. Spheroid culture as a tool for creating 3D complex tissues. Trends Biotechnol 2013;31:108‐115. - PubMed
-
- Luni C, Serena E, Elvassore N. Human‐on‐chip for therapy development and fundamental science. Curr Opin Biotechnol 2014;25:45‐50. - PubMed

