Baculoviral delivery of CRISPR/Cas9 facilitates efficient genome editing in human cells
- PMID: 28640891
- PMCID: PMC5480884
- DOI: 10.1371/journal.pone.0179514
Baculoviral delivery of CRISPR/Cas9 facilitates efficient genome editing in human cells
Abstract
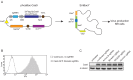
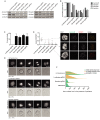
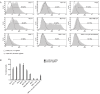
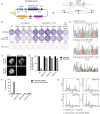
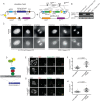
The CRISPR/Cas9 system is a highly effective tool for genome editing. Key to robust genome editing is the efficient delivery of the CRISPR/Cas9 machinery. Viral delivery systems are efficient vehicles for the transduction of foreign genes but commonly used viral vectors suffer from a limited capacity in the genetic information they can carry. Baculovirus however is capable of carrying large exogenous DNA fragments. Here we investigate the use of baculoviral vectors as a delivery vehicle for CRISPR/Cas9 based genome-editing tools. We demonstrate transduction of a panel of cell lines with Cas9 and an sgRNA sequence, which results in efficient knockout of all four targeted subunits of the chromosomal passenger complex (CPC). We further show that introduction of a homology directed repair template into the same CRISPR/Cas9 baculovirus facilitates introduction of specific point mutations and endogenous gene tags. Tagging of the CPC recruitment factor Haspin with the fluorescent reporter YFP allowed us to study its native localization as well as recruitment to the cohesin subunit Pds5B.
Conflict of interest statement
Figures
Similar articles
-
CRISPR-Cas9 vectors for genome editing and host engineering in the baculovirus-insect cell system.Proc Natl Acad Sci U S A. 2017 Aug 22;114(34):9068-9073. doi: 10.1073/pnas.1705836114. Epub 2017 Aug 7. Proc Natl Acad Sci U S A. 2017. PMID: 28784806 Free PMC article.
-
Improvement of baculovirus as protein expression vector and as biopesticide by CRISPR/Cas9 editing.Biotechnol Bioeng. 2019 Nov;116(11):2823-2833. doi: 10.1002/bit.27139. Epub 2019 Sep 2. Biotechnol Bioeng. 2019. PMID: 31403180
-
Comparison of CRISPR-Cas9 Tools for Transcriptional Repression and Gene Disruption in the BEVS.Viruses. 2021 Sep 24;13(10):1925. doi: 10.3390/v13101925. Viruses. 2021. PMID: 34696355 Free PMC article.
-
Delivery of CRISPR/Cas9 for therapeutic genome editing.J Gene Med. 2019 Jul;21(7):e3107. doi: 10.1002/jgm.3107. J Gene Med. 2019. PMID: 31237055 Review.
-
CRISPR/Cas9-based epigenome editing: An overview of dCas9-based tools with special emphasis on off-target activity.Methods. 2019 Jul 15;164-165:109-119. doi: 10.1016/j.ymeth.2019.05.003. Epub 2019 May 6. Methods. 2019. PMID: 31071448 Review.
Cited by
-
Untangling the contribution of Haspin and Bub1 to Aurora B function during mitosis.J Cell Biol. 2020 Mar 2;219(3):e201907087. doi: 10.1083/jcb.201907087. J Cell Biol. 2020. PMID: 32027339 Free PMC article.
-
Afadin loss induces breast cancer metastasis through destabilisation of E-cadherin to F-actin linkage.J Pathol. 2025 May;266(1):26-39. doi: 10.1002/path.6394. Epub 2025 Mar 3. J Pathol. 2025. PMID: 40026293 Free PMC article.
-
Human chromosome-specific aneuploidy is influenced by DNA-dependent centromeric features.EMBO J. 2020 Jan 15;39(2):e102924. doi: 10.15252/embj.2019102924. Epub 2019 Nov 21. EMBO J. 2020. PMID: 31750958 Free PMC article.
-
Highly efficient CRISPR-mediated large DNA docking and multiplexed prime editing using a single baculovirus.Nucleic Acids Res. 2022 Jul 22;50(13):7783-7799. doi: 10.1093/nar/gkac587. Nucleic Acids Res. 2022. PMID: 35801912 Free PMC article.
-
The Ins and Outs of Aurora B Inner Centromere Localization.Front Cell Dev Biol. 2017 Dec 22;5:112. doi: 10.3389/fcell.2017.00112. eCollection 2017. Front Cell Dev Biol. 2017. PMID: 29312936 Free PMC article. Review.
References
-
- Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–23. doi: 10.1126/science.1231143 ; PubMed Central PMCID: PMCPMC3795411. - DOI - PMC - PubMed
-
- Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature. 2015;520(7546):186–91. doi: 10.1038/nature14299 ; PubMed Central PMCID: PMCPMC4393360. - DOI - PMC - PubMed
-
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816–21. doi: 10.1126/science.1225829 . - DOI - PMC - PubMed
-
- Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339(6121):823–6. doi: 10.1126/science.1232033 ; PubMed Central PMCID: PMCPMC3712628. - DOI - PMC - PubMed
-
- Mao Z, Bozzella M, Seluanov A, Gorbunova V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair (Amst). 2008;7(10):1765–71. doi: 10.1016/j.dnarep.2008.06.018 ; PubMed Central PMCID: PMCPMC2695993. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

