Hierarchical cortical transcriptome disorganization in autism
- PMID: 28649314
- PMCID: PMC5480153
- DOI: 10.1186/s13229-017-0147-7
Hierarchical cortical transcriptome disorganization in autism
Abstract
Background: Autism spectrum disorders (ASD) are etiologically heterogeneous and complex. Functional genomics work has begun to identify a diverse array of dysregulated transcriptomic programs (e.g., synaptic, immune, cell cycle, DNA damage, WNT signaling, cortical patterning and differentiation) potentially involved in ASD brain abnormalities during childhood and adulthood. However, it remains unclear whether such diverse dysregulated pathways are independent of each other or instead reflect coordinated hierarchical systems-level pathology.
Methods: Two ASD cortical transcriptome datasets were re-analyzed using consensus weighted gene co-expression network analysis (WGCNA) to identify common co-expression modules across datasets. Linear mixed-effect models and Bayesian replication statistics were used to identify replicable differentially expressed modules. Eigengene network analysis was then utilized to identify between-group differences in how co-expression modules interact and cluster into hierarchical meta-modular organization. Protein-protein interaction analyses were also used to determine whether dysregulated co-expression modules show enhanced interactions.
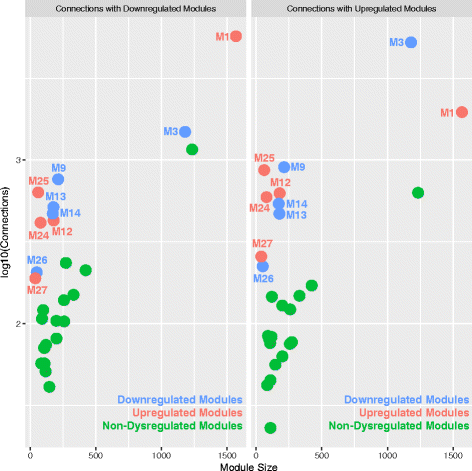
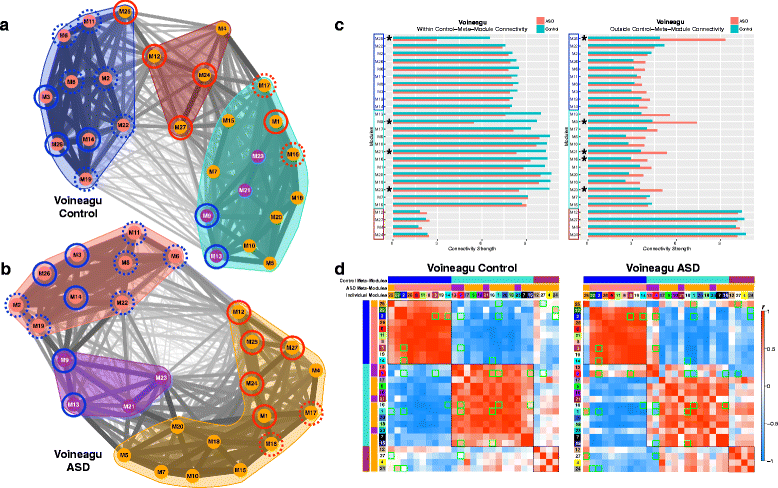
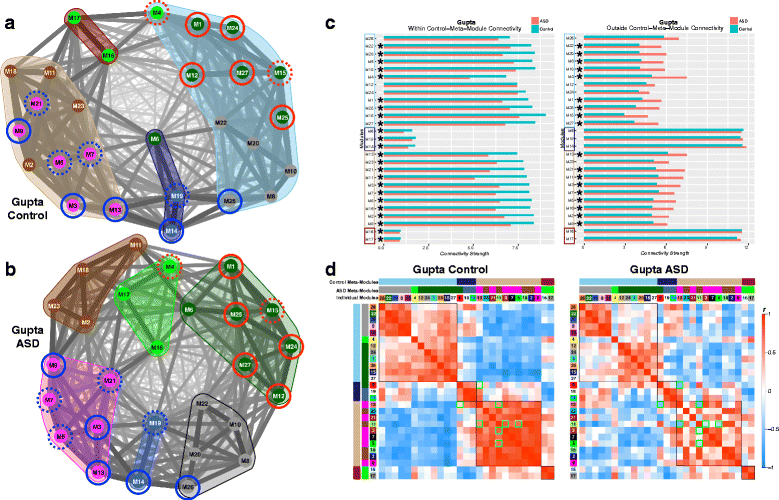
Results: We find replicable evidence for 10 gene co-expression modules that are differentially expressed in ASD cortex. Rather than being independent non-interacting sources of pathology, these dysregulated co-expression modules work in synergy and physically interact at the protein level. These systems-level transcriptional signals are characterized by downregulation of synaptic processes coordinated with upregulation of immune/inflammation, response to other organism, catabolism, viral processes, translation, protein targeting and localization, cell proliferation, and vasculature development. Hierarchical organization of meta-modules (clusters of highly correlated modules) is also highly affected in ASD.
Conclusions: These findings highlight that dysregulation of the ASD cortical transcriptome is characterized by the dysregulation of multiple coordinated transcriptional programs producing synergistic systems-level effects that cannot be fully appreciated by studying the individual component biological processes in isolation.
Keywords: Autism; Gene co-expression networks; Immune; Synapse; Systems biology; Transcriptome; Translation.
Figures





References
-
- Chow ML, Pramparo T, Winn ME, Barnes CC, Li HR, Weiss L, Fan JB, Murray S, April C, Belinson H, et al. Age-dependent brain gene expression and copy number anomalies in autism suggest distinct pathological processes at young versus mature ages. PLoS Genet. 2012;8:e1002592. doi: 10.1371/journal.pgen.1002592. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

