A Massively Parallel Fluorescence Assay to Characterize the Effects of Synonymous Mutations on TP53 Expression
- PMID: 28652265
- PMCID: PMC5626615
- DOI: 10.1158/1541-7786.MCR-17-0245
A Massively Parallel Fluorescence Assay to Characterize the Effects of Synonymous Mutations on TP53 Expression
Abstract
Although synonymous mutations can affect gene expression, they have generally not been considered in genomic studies that focus on mutations that increase the risk of cancer. However, mounting evidence implicates some synonymous mutations as driver mutations in cancer. Here, a massively parallel assay, based on cell sorting of a reporter containing a segment of p53 fused to GFP, was used to measure the effects of nearly all synonymous mutations in exon 6 of TP53 In this reporter context, several mutations within the exon caused strong expression changes including mutations that may cause potential gain or loss of function. Further analysis indicates that these effects are largely attributed to errors in splicing, including exon skipping, intron inclusion, and exon truncation, resulting from mutations both at exon-intron junctions and within the body of the exon. These mutations are found at extremely low frequencies in healthy populations and are enriched a few-fold in cancer genomes, suggesting that some of them may be driver mutations in TP53 This assay provides a general framework to identify previously unknown detrimental synonymous mutations in cancer genes.Implications: Using a massively parallel assay, this study demonstrates that synonymous mutations in the TP53 gene affect protein expression, largely through their impact on splicing.Visual Overview: http://mcr.aacrjournals.org/content/molcanres/15/10/1301/F1.large.jpg Mol Cancer Res; 15(10); 1301-7. ©2017 AACR.
©2017 American Association for Cancer Research.
Conflict of interest statement
The authors have no conflicts of interest.
Figures

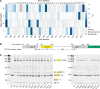

References
-
- Supek F, Miñana B, Valcárcel J, Gabaldón T, Lehner B. Synonymous mutations frequently act as driver mutations in human cancers. Cell. 2014;156:1324–35. - PubMed
-
- Sauna ZE, Kimchi-Sarfaty C. Understanding the contribution of synonymous mutations to human disease. Nat Rev Genet. 2011;12:683–91. - PubMed
-
- Cartegni L, Chew SL, Krainer AR. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Vol. 3. Nature Publishing Group; 2002. pp. 285–98. - PubMed
-
- Brest P, Lapaquette P, Souidi M, Lebrigand K, Cesaro A, Vouret-Craviari V, et al. A synonymous variant in IRGM alters a binding site for miR-196 and causes deregulation of IRGM-dependent xenophagy in Crohn's disease. Nat Genet. 2011;43:242–5. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

