Stabilization of a nucleotide-binding domain of the cystic fibrosis transmembrane conductance regulator yields insight into disease-causing mutations
- PMID: 28655774
- PMCID: PMC5572908
- DOI: 10.1074/jbc.M116.772335
Stabilization of a nucleotide-binding domain of the cystic fibrosis transmembrane conductance regulator yields insight into disease-causing mutations
Abstract
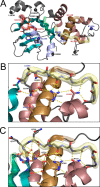
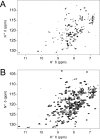
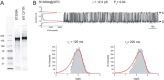
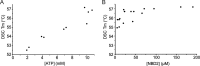
Characterization of the second nucleotide-binding domain (NBD2) of the cystic fibrosis transmembrane conductance regulator (CFTR) has lagged behind research into the NBD1 domain, in part because NBD1 contains the F508del mutation, which is the dominant cause of cystic fibrosis. Research on NBD2 has also been hampered by the overall instability of the domain and the difficulty of producing reagents. Nonetheless, multiple disease-causing mutations reside in NBD2, and the domain is critical for CFTR function, because channel gating involves NBD1/NBD2 dimerization, and NBD2 contains the catalytically active ATPase site in CFTR. Recognizing the paucity of structural and biophysical data on NBD2, here we have defined a bioinformatics-based method for manually identifying stabilizing substitutions in NBD2, and we used an iterative process of screening single substitutions against thermal melting points to both produce minimally mutated stable constructs and individually characterize mutations. We present a range of stable constructs with minimal mutations to help inform further research on NBD2. We have used this stabilized background to study the effects of NBD2 mutations identified in cystic fibrosis (CF) patients, demonstrating that mutants such as N1303K and G1349D are characterized by lower stability, as shown previously for some NBD1 mutations, suggesting a potential role for NBD2 instability in the pathology of CF.
Keywords: ABC transporter; cystic fibrosis transmembrane conductance regulator (CFTR); protein engineering; protein misfolding; protein stability.
© 2017 by The American Society for Biochemistry and Molecular Biology, Inc.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures




References
-
- Riordan J. R., Rommens J. M., Kerem B., Alon N., Rozmahel R., Grzelczak Z., Zielenski J., Lok S., Plavsic N., and Chou J. L. (1989) Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 245, 1066–1073 - PubMed
-
- Hyde S. C., Emsley P., Hartshorn M. J., Mimmack M. M., Gileadi U., Pearce S. R., Gallagher M. P., Gill D. R., Hubbard R. E., and Higgins C. F. (1990) Structural model of ATP-binding proteins associated with cystic fibrosis, multidrug resistance and bacterial transport. Nature 346, 362–365 - PubMed
-
- Anderson M. P., Gregory R. J., Thompson S., Souza D. W., Paul S., Mulligan R. C., Smith A. E., and Welsh M. J. (1991) Demonstration that CFTR is a chloride channel by alteration of its anion selectivity. Science 253, 202–205 - PubMed
-
- Winter M. C., and Welsh M. J. (1997) Stimulation of CFTR activity by its phosphorylated R domain. Nature 389, 294–296 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

