Hepatocyte Hyperproliferation upon Liver-Specific Co-disruption of Thioredoxin-1, Thioredoxin Reductase-1, and Glutathione Reductase
- PMID: 28658624
- PMCID: PMC5730093
- DOI: 10.1016/j.celrep.2017.06.019
Hepatocyte Hyperproliferation upon Liver-Specific Co-disruption of Thioredoxin-1, Thioredoxin Reductase-1, and Glutathione Reductase
Abstract
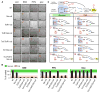
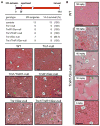
Energetic nutrients are oxidized to sustain high intracellular NADPH/NADP+ ratios. NADPH-dependent reduction of thioredoxin-1 (Trx1) disulfide and glutathione disulfide by thioredoxin reductase-1 (TrxR1) and glutathione reductase (Gsr), respectively, fuels antioxidant systems and deoxyribonucleotide synthesis. Mouse livers lacking both TrxR1 and Gsr sustain these essential activities using an NADPH-independent methionine-consuming pathway; however, it remains unclear how this reducing power is distributed. Here, we show that liver-specific co-disruption of the genes encoding Trx1, TrxR1, and Gsr (triple-null) causes dramatic hepatocyte hyperproliferation. Thus, even in the absence of Trx1, methionine-fueled glutathione production supports hepatocyte S phase deoxyribonucleotide production. Also, Trx1 in the absence of TrxR1 provides a survival advantage to cells under hyperglycemic stress, suggesting that glutathione, likely via glutaredoxins, can reduce Trx1 disulfide in vivo. In triple-null livers like in many cancers, deoxyribonucleotide synthesis places a critical yet relatively low-volume demand on these reductase systems, thereby favoring high hepatocyte turnover over sustained hepatocyte integrity.
Keywords: cancer; glutathione; liver; methionine cycle; mouse model; proliferation; redox; ribonucleotide reductase; thioredoxin; transsulfuration.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures




Similar articles
-
TrxR1, Gsr, and oxidative stress determine hepatocellular carcinoma malignancy.Proc Natl Acad Sci U S A. 2019 Jun 4;116(23):11408-11417. doi: 10.1073/pnas.1903244116. Epub 2019 May 16. Proc Natl Acad Sci U S A. 2019. PMID: 31097586 Free PMC article.
-
Dietary methionine can sustain cytosolic redox homeostasis in the mouse liver.Nat Commun. 2015 Mar 20;6:6479. doi: 10.1038/ncomms7479. Nat Commun. 2015. PMID: 25790857 Free PMC article.
-
Glutathione and glutaredoxin act as a backup of human thioredoxin reductase 1 to reduce thioredoxin 1 preventing cell death by aurothioglucose.J Biol Chem. 2012 Nov 2;287(45):38210-9. doi: 10.1074/jbc.M112.392225. Epub 2012 Sep 13. J Biol Chem. 2012. PMID: 22977247 Free PMC article.
-
Interplay between cytosolic disulfide reductase systems and the Nrf2/Keap1 pathway.Biochem Soc Trans. 2015 Aug;43(4):632-8. doi: 10.1042/BST20150021. Epub 2015 Aug 3. Biochem Soc Trans. 2015. PMID: 26551704 Free PMC article. Review.
-
The thioredoxin antioxidant system.Free Radic Biol Med. 2014 Jan;66:75-87. doi: 10.1016/j.freeradbiomed.2013.07.036. Epub 2013 Jul 27. Free Radic Biol Med. 2014. PMID: 23899494 Review.
Cited by
-
Reactive Oxygen Species-Induced Lipid Peroxidation in Apoptosis, Autophagy, and Ferroptosis.Oxid Med Cell Longev. 2019 Oct 13;2019:5080843. doi: 10.1155/2019/5080843. eCollection 2019. Oxid Med Cell Longev. 2019. PMID: 31737171 Free PMC article. Review.
-
Redox Regulation via Glutaredoxin-1 and Protein S-Glutathionylation.Antioxid Redox Signal. 2020 Apr 1;32(10):677-700. doi: 10.1089/ars.2019.7963. Epub 2020 Jan 23. Antioxid Redox Signal. 2020. PMID: 31813265 Free PMC article. Review.
-
Thioredoxin-1 maintains mitochondrial function via mechanistic target of rapamycin signalling in the heart.Cardiovasc Res. 2020 Aug 1;116(10):1742-1755. doi: 10.1093/cvr/cvz251. Cardiovasc Res. 2020. PMID: 31584633 Free PMC article.
-
Glutathione-Related Enzymes and Proteins: A Review.Molecules. 2023 Feb 2;28(3):1447. doi: 10.3390/molecules28031447. Molecules. 2023. PMID: 36771108 Free PMC article. Review.
-
Emerging Evidence of the Significance of Thioredoxin-1 in Hematopoietic Stem Cell Aging.Antioxidants (Basel). 2022 Jun 29;11(7):1291. doi: 10.3390/antiox11071291. Antioxidants (Basel). 2022. PMID: 35883782 Free PMC article. Review.
References
-
- Arnér ESJ. Focus on mammalian thioredoxin reductases–important selenoproteins with versatile functions. Biochim Biophys Acta. 2009;1790:495–526. - PubMed
-
- Arnér ESJ, Eriksson S. Mammalian deoxyribonucleoside kinases. Pharmacol Ther. 1995;67:155–186. - PubMed
-
- Arnér ES, Holmgren A. Physiological functions of thioredoxin and thioredoxin reductase. Eur J Biochem. 2000;267:6102–6109. - PubMed
-
- Brand MD, Affourtit C, Esteves TC, Green K, Lambert AJ, Miwa S, Pakay JL, Parker N. Mitochondrial superoxide: production, biological effects, and activation of uncoupling proteins. Free Radic Biol Med. 2004;37:755–767. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

