Monophyletic Origin and Evolution of the Largest Crucifer Genomes
- PMID: 28667048
- PMCID: PMC5543974
- DOI: 10.1104/pp.17.00457
Monophyletic Origin and Evolution of the Largest Crucifer Genomes
Abstract
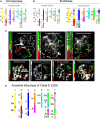
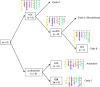
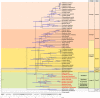
Clade E, or the Hesperis clade, is one of the major Brassicaceae (Crucifereae) clades, comprising some 48 genera and 351 species classified into seven tribes and is distributed predominantly across arid and montane regions of Asia. Several taxa have socioeconomic significance, being important ornamental but also weedy and invasive species. From the comparative genomic perspective, the clade is noteworthy as it harbors species with the largest crucifer genomes but low numbers of chromosomes (n = 5-7). By applying comparative cytogenetic analysis and whole-chloroplast phylogenetics, we constructed, to our knowledge, the first partial and complete cytogenetic maps for selected representatives of clade E tribes and investigated their relationships in a family-wide context. The Hesperis clade is a well-supported monophyletic lineage comprising seven tribes: Anchonieae, Buniadeae, Chorisporeae, Dontostemoneae, Euclidieae, Hesperideae, and Shehbazieae. The clade diverged from other Brassicaceae crown-group clades during the Oligocene, followed by subsequent Miocene tribal diversifications in central/southwestern Asia. The inferred ancestral karyotype of clade E (CEK; n = 7) originated from an older n = 8 genome, which also was the purported progenitor of tribe Arabideae (KAA genome). In most taxa of clade E, the seven linkage groups of CEK either remained conserved (Chorisporeae) or were reshuffled by chromosomal translocations (Euclidieae). In 50% of Anchonieae and Hesperideae species, the CEK genome has undergone descending dysploidy toward n = 6 (-5). These genomic data elucidate early genome evolution in Brassicaceae and pave the way for future whole-genome sequencing and assembly efforts in this as yet genomically neglected group of crucifer plants.
© 2017 American Society of Plant Biologists. All Rights Reserved.
Figures

Similar articles
-
The large genome size variation in the Hesperis clade was shaped by the prevalent proliferation of DNA repeats and rarer genome downsizing.Ann Bot. 2019 Aug 2;124(1):103-120. doi: 10.1093/aob/mcz036. Ann Bot. 2019. PMID: 31220201 Free PMC article.
-
Chromosomal phylogeny and karyotype evolution in x=7 crucifer species (Brassicaceae).Plant Cell. 2008 Oct;20(10):2559-70. doi: 10.1105/tpc.108.062166. Epub 2008 Oct 3. Plant Cell. 2008. PMID: 18836039 Free PMC article.
-
Linked by Ancestral Bonds: Multiple Whole-Genome Duplications and Reticulate Evolution in a Brassicaceae Tribe.Mol Biol Evol. 2021 May 4;38(5):1695-1714. doi: 10.1093/molbev/msaa327. Mol Biol Evol. 2021. PMID: 33331908 Free PMC article.
-
The ABC's of comparative genomics in the Brassicaceae: building blocks of crucifer genomes.Trends Plant Sci. 2006 Nov;11(11):535-42. doi: 10.1016/j.tplants.2006.09.002. Epub 2006 Oct 6. Trends Plant Sci. 2006. PMID: 17029932 Review.
-
Comparative paleogenomics of crucifers: ancestral genomic blocks revisited.Curr Opin Plant Biol. 2016 Apr;30:108-15. doi: 10.1016/j.pbi.2016.02.001. Epub 2016 Mar 4. Curr Opin Plant Biol. 2016. PMID: 26945766 Review.
Cited by
-
Genomic Blocks in Aethionema arabicum Support Arabideae as Next Diverging Clade in Brassicaceae.Front Plant Sci. 2020 Jun 3;11:719. doi: 10.3389/fpls.2020.00719. eCollection 2020. Front Plant Sci. 2020. PMID: 32582250 Free PMC article.
-
Nested whole-genome duplications coincide with diversification and high morphological disparity in Brassicaceae.Nat Commun. 2020 Jul 30;11(1):3795. doi: 10.1038/s41467-020-17605-7. Nat Commun. 2020. PMID: 32732942 Free PMC article.
-
The genome of Orychophragmus violaceus provides genomic insights into the evolution of Brassicaceae polyploidization and its distinct traits.Plant Commun. 2023 Mar 13;4(2):100431. doi: 10.1016/j.xplc.2022.100431. Epub 2022 Sep 7. Plant Commun. 2023. PMID: 36071668 Free PMC article.
-
Genome Evolution in Arabideae Was Marked by Frequent Centromere Repositioning.Plant Cell. 2020 Mar;32(3):650-665. doi: 10.1105/tpc.19.00557. Epub 2020 Jan 9. Plant Cell. 2020. PMID: 31919297 Free PMC article.
-
Celebrating Mendel, McClintock, and Darlington: On end-to-end chromosome fusions and nested chromosome fusions.Plant Cell. 2022 Jul 4;34(7):2475-2491. doi: 10.1093/plcell/koac116. Plant Cell. 2022. PMID: 35441689 Free PMC article. Review.
References
-
- Al-Shehbaz IA. (2012) A generic and tribal synopsis of the Brassicaceae (Cruciferae). Taxon 61: 931–954
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215: 403–410 - PubMed
-
- Beilstein MA, Al-Shehbaz IA, Kellogg EA (2006) Brassicaceae phylogeny and trichome evolution. Am J Bot 93: 607–619 - PubMed
-
- Beilstein MA, Al-Shehbaz IA, Mathews S, Kellogg EA (2008) Brassicaceae phylogeny inferred from phytochrome A and ndhF sequence data: tribes and trichomes revisited. Am J Bot 95: 1307–1327 - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

