Methods to Monitor and Quantify Autophagy in the Social Amoeba Dictyostelium discoideum
- PMID: 28671610
- PMCID: PMC5617964
- DOI: 10.3390/cells6030018
Methods to Monitor and Quantify Autophagy in the Social Amoeba Dictyostelium discoideum
Abstract
Autophagy is a eukaryotic catabolic pathway that degrades and recycles cellular components to maintain homeostasis. It can target protein aggregates, superfluous biomolecular complexes, dysfunctional and damaged organelles, as well as pathogenic intracellular microbes. Autophagy is a dynamic process in which the different stages from initiation to final degradation of cargo are finely regulated. Therefore, the study of this process requires the use of a palette of techniques, which are continuously evolving and whose interpretation is not trivial. Here, we present the social amoeba Dictyostelium discoideum as a relevant model to study autophagy. Several methods have been developed based on the tracking and observation of autophagosomes by microscopy, analysis of changes in expression of autophagy genes and proteins, and examination of the autophagic flux with various techniques. In this review, we discuss the pros and cons of the currently available techniques to assess autophagy in this organism.
Keywords: Dictyostelium; autophagic markers; autophagy; cleavage assays; flux assays.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






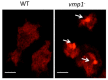
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

