HHV-6A/B Integration and the Pathogenesis Associated with the Reactivation of Chromosomally Integrated HHV-6A/B
- PMID: 28672870
- PMCID: PMC5537652
- DOI: 10.3390/v9070160
HHV-6A/B Integration and the Pathogenesis Associated with the Reactivation of Chromosomally Integrated HHV-6A/B
Abstract
Unlike other human herpesviruses, human herpesvirus 6A and 6B (HHV-6A/B) infection can lead to integration of the viral genome in human chromosomes. When integration occurs in germinal cells, the integrated HHV-6A/B genome can be transmitted to 50% of descendants. Such individuals, carrying one copy of the HHV-6A/B genome in every cell, are referred to as having inherited chromosomally-integrated HHV-6A/B (iciHHV-6) and represent approximately 1% of the world's population. Interestingly, HHV-6A/B integrate their genomes in a specific region of the chromosomes known as telomeres. Telomeres are located at chromosomes' ends and play essential roles in chromosomal stability and the long-term proliferative potential of cells. Considering that the integrated HHV-6A/B genome is mostly intact without any gross rearrangements or deletions, integration is likely used for viral maintenance into host cells. Knowing the roles played by telomeres in cellular homeostasis, viral integration in such structure is not likely to be without consequences. At present, the mechanisms and factors involved in HHV-6A/B integration remain poorly defined. In this review, we detail the potential biological and medical impacts of HHV-6A/B integration as well as the possible chromosomal integration and viral excision processes.
Keywords: human herpesvirus 6A/B; inherited chromosomally-integrated HHV-6A/B; integration; reactivation; telomere.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

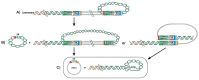
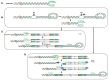
References
-
- Salahuddin S.Z., Ablashi D.V., Markham P.D., Josephs S.F., Sturzenegger S., Kaplan M., Halligan G., Biberfeld P., Wong-Staal F., Kramarsky B., et al. Isolation of a new virus, HBLV, in patients with lymphoproliferative disorders. Science. 1986;234:596–601. doi: 10.1126/science.2876520. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

