Pseudomonas syringae pv. tomato DC3000 polymutants deploying coronatine and two type III effectors produce quantifiable chlorotic spots from individual bacterial colonies in Nicotiana benthamiana leaves
- PMID: 28677296
- PMCID: PMC6637995
- DOI: 10.1111/mpp.12579
Pseudomonas syringae pv. tomato DC3000 polymutants deploying coronatine and two type III effectors produce quantifiable chlorotic spots from individual bacterial colonies in Nicotiana benthamiana leaves
Abstract
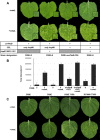
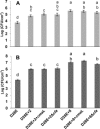
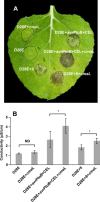
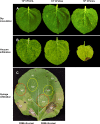
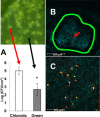
Primary virulence factors of Pseudomonas syringae pv. tomato DC3000 include the phytotoxin coronatine (COR) and a repertoire of 29 effector proteins injected into plant cells by the type III secretion system (T3SS). DC3000 derivatives differentially producing COR, the T3SS machinery and subsets of key effectors were constructed and assayed in leaves of Nicotiana benthamiana. Bacteria were inoculated by the dipping of whole plants and assayed for population growth and the production of chlorotic spots on leaves. The strains fell into three classes. Class I strains are T3SS+ but functionally effectorless, grow poorly in planta and produce faint chlorotic spots only if COR+ . Class II strains are T3SS- or, if T3SS+ , also produce effectors AvrPtoB and HopM1. Class II strains grow better than class I strains in planta and, if COR+ , produce robust chlorotic spots. Class III strains are T3SS+ and minimally produce AvrPtoB, HopM1 and three other effectors encoded in the P. syringae conserved effector locus. These strains differ from class II strains in growing better in planta, and produce chlorotic spots without COR if the precursor coronafacic acid is produced. Assays for chlorotic spot formation, in conjunction with pressure infiltration of low-level inoculum and confocal microscopy of fluorescent protein-labelled bacteria, revealed that single bacteria in the apoplast are capable of producing colonies and associated leaf spots in a 1 : 1 : 1 manner. However, COR makes no significant contribution to the bacterial colonization of the apoplast, but, instead, enables a gratuitous, semi-quantitative, surface indicator of bacterial growth, which is determined by the strain's effector composition.
Keywords: bacterial plant colonization; bacterial polymutants; coronafacic acid; coronatine; effectors; type III secretion system; virulence assays.
© 2017 BSPP AND JOHN WILEY & SONS LTD.
Figures

Similar articles
-
Defining essential processes in plant pathogenesis with Pseudomonas syringae pv. tomato DC3000 disarmed polymutants and a subset of key type III effectors.Mol Plant Pathol. 2018 Jul;19(7):1779-1794. doi: 10.1111/mpp.12655. Epub 2018 Feb 1. Mol Plant Pathol. 2018. PMID: 29277959 Free PMC article. Review.
-
CorR regulates multiple components of virulence in Pseudomonas syringae pv. tomato DC3000.Mol Plant Microbe Interact. 2006 Jul;19(7):768-79. doi: 10.1094/MPMI-19-0768. Mol Plant Microbe Interact. 2006. PMID: 16838789
-
A Pseudomonas syringae pv. tomato avrE1/hopM1 mutant is severely reduced in growth and lesion formation in tomato.Mol Plant Microbe Interact. 2006 Feb;19(2):99-111. doi: 10.1094/MPMI-19-0099. Mol Plant Microbe Interact. 2006. PMID: 16529372
-
Pseudomonas syringae pv. tomato DC3000 CmaL (PSPTO4723), a DUF1330 family member, is needed to produce L-allo-isoleucine, a precursor for the phytotoxin coronatine.J Bacteriol. 2013 Jan;195(2):287-96. doi: 10.1128/JB.01352-12. Epub 2012 Nov 9. J Bacteriol. 2013. PMID: 23144243 Free PMC article.
-
The phytotoxin coronatine is a multifunctional component of the virulence armament of Pseudomonas syringae.Planta. 2014 Dec;240(6):1149-65. doi: 10.1007/s00425-014-2151-x. Epub 2014 Aug 26. Planta. 2014. PMID: 25156488 Free PMC article. Review.
Cited by
-
Pseudomonas syringae AlgU Downregulates Flagellin Gene Expression, Helping Evade Plant Immunity.J Bacteriol. 2020 Jan 29;202(4):e00418-19. doi: 10.1128/JB.00418-19. Print 2020 Jan 29. J Bacteriol. 2020. PMID: 31740494 Free PMC article.
-
Ca2+-Induced Two-Component System CvsSR Regulates the Type III Secretion System and the Extracytoplasmic Function Sigma Factor AlgU in Pseudomonas syringae pv. tomato DC3000.J Bacteriol. 2018 Feb 7;200(5):e00538-17. doi: 10.1128/JB.00538-17. Print 2018 Mar 1. J Bacteriol. 2018. PMID: 29263098 Free PMC article.
-
Genetic dissection of the tissue-specific roles of type III effectors and phytotoxins in the pathogenicity of Pseudomonas syringae pv. syringae to cherry.Mol Plant Pathol. 2024 Apr;25(4):e13451. doi: 10.1111/mpp.13451. Mol Plant Pathol. 2024. PMID: 38590135 Free PMC article.
-
Draft Genome Sequences of Pseudomonas syringae pv. tomato Strains J4 and J6, Isolated in Florida.Microbiol Resour Announc. 2021 Apr 15;10(15):e00127-21. doi: 10.1128/MRA.00127-21. Microbiol Resour Announc. 2021. PMID: 33858923 Free PMC article.
-
Cooperative virulence via the collective action of secreted pathogen effectors.Nat Microbiol. 2023 Apr;8(4):640-650. doi: 10.1038/s41564-023-01328-8. Epub 2023 Feb 13. Nat Microbiol. 2023. PMID: 36782026
References
-
- Badel, J.L. , Nomura, K. , Bandyopadhyay, S. , Shimizu, R. , Collmer, A. and He, S.Y. (2003) Pseudomonas syringae pv. tomato DC3000 HopPtoM (CEL ORF3) is important for lesion formation but not growth in tomato and is secreted and translocated by the Hrp type III secretion system in a chaperone‐dependent manner. Mol. Microbiol. 49, 1239–1251. - PubMed
-
- Badel, J.L. , Shimizu, R. , Oh, H.‐S. and Collmer, A. (2006) A Pseudomonas syringae pv. tomato avrE1/hopM1 mutant is severely reduced in growth and lesion formation in tomato. Mol. Plant–Microbe Interact. 19, 99–111. - PubMed
-
- Baltrus, D.A. , Nishimura, M.T. , Romanchuk, A. , Chang, J.H. , Mukhtar, M.S. , Cherkis, K. , Roach, J. , Grant, S.R. , Jones, C.D. and Dangl, J.L. (2011) Dynamic evolution of pathogenicity revealed by sequencing and comparative genomics of 19 Pseudomonas syringae isolates. PLoS Pathog. 7, e1002132. - PMC - PubMed
-
- Bombarely, A. , Rosli, H.G. , Vrebalov, J. , Moffett, P. , Mueller, L.A. and Martin, G.B. (2012) A draft genome sequence of Nicotiana benthamiana to enhance molecular plant–microbe biology research. Mol. Plant–Microbe Interact. 12, 1523–1530. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

