Three members of Medicago truncatula ST family are ubiquitous during development and modulated by nutritional status (MtST1) and dehydration (MtST2 and MtST3)
- PMID: 28693485
- PMCID: PMC5504553
- DOI: 10.1186/s12870-017-1061-z
Three members of Medicago truncatula ST family are ubiquitous during development and modulated by nutritional status (MtST1) and dehydration (MtST2 and MtST3)
Abstract
Background: ShooT specific/Specific Tissue (ST) belong to a protein family of unknown function characterized by the DUF2775 domain and produced in specific taxonomic plant families, mainly Fabaceae and Asteraceae, with the Medicago truncatula ST family being the largest. The putative roles proposed for this family are cell elongation, biotic interactions, abiotic stress and N reserve. The aim of this work was to go deeper into the role of three M. truncatula ST proteins, namely ST1, ST2 and ST3. Our starting hypothesis was that each member of the family could perform a specific role, and hence, each ST gene would be subjected to a different type of regulation.
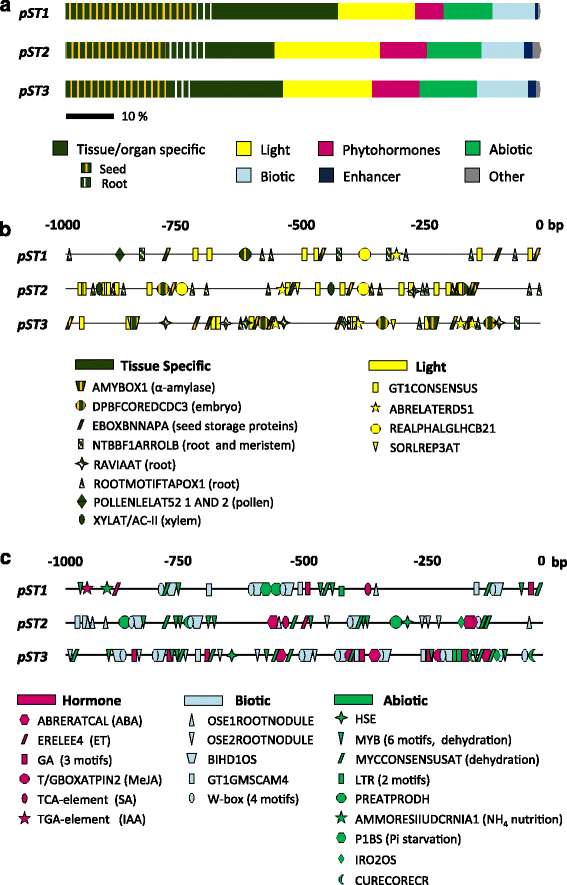
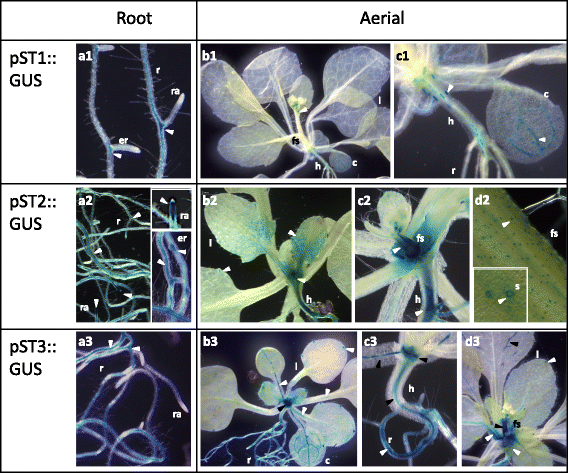
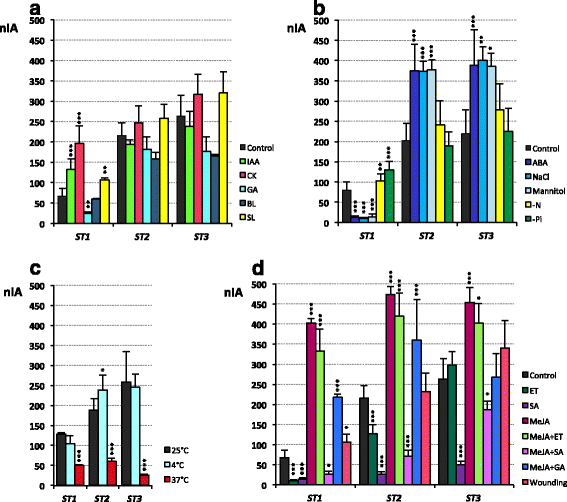
Results: The search for cis-acting regulatory elements (CREs) in silico in pST1, pST2 and pST3 promoters showed prevalence of tissue/organ specific motifs, especially root- and seed-specific ones. Light, hormone, biotic and abiotic related motifs were also present. None of these pSTs showed the same combination of CREs, or presented the same activity pattern. In general, pST activity was associated with the vascular cylinder, mainly in roots. Promoter activation was highly specific and dissimilar during reproductive development. The ST1, ST2 and ST3 transcripts accumulated in most of the organs and developmental stages analysed - decreasing with age - and expression was higher in the roots than in the aerial parts and more abundant in light-grown plants. The effect of the different treatments on transcript accumulation indicated that ST1 behaved differently from ST2 and ST3, mainly in response to several hormones and dehydration treatments (NaCl or mannitol), upon which ST1 transcript levels decreased and ST2 and ST3 levels increased. Finally, the ST1 protein was located in the cell wall whereas ST2 and ST3 were present both in the cytoplasm and in the cell wall.
Conclusions: The ST proteins studied are ubiquitous proteins that could perform distinct/complementary roles in plant biology as they are encoded by differentially regulated genes. Based on these differences we have established two functional groups among the three STs. ST1 would participate in processes affected by nutritional status, while ST2 and ST3 seem to act when plants are challenged with abiotic stresses related to water stress and in physiologically controlled desiccation processes such as the seed maturation.
Keywords: Abiotic stress; DUF2775; Development; Medicago truncatula; ST protein; cis-acting regulatory elements.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures




Similar articles
-
Specific tissue proteins 1 and 6 are involved in root biology during normal development and under symbiotic and pathogenic interactions in Medicago truncatula.Planta. 2021 Jan 2;253(1):7. doi: 10.1007/s00425-020-03538-4. Planta. 2021. PMID: 33387090
-
A genome-wide study of the lipoxygenase gene families in Medicago truncatula and Medicago sativa reveals that MtLOX24 participates in the methyl jasmonate response.BMC Genomics. 2024 Feb 19;25(1):195. doi: 10.1186/s12864-024-10071-1. BMC Genomics. 2024. PMID: 38373903 Free PMC article.
-
Three members of Medicago truncatula ST family (MtST4, MtST5 and MtST6) are specifically induced by hormones involved in biotic interactions.Plant Physiol Biochem. 2018 Jun;127:496-505. doi: 10.1016/j.plaphy.2018.04.019. Epub 2018 Apr 21. Plant Physiol Biochem. 2018. PMID: 29705570
-
In Medicago truncatula, water deficit modulates the transcript accumulation of components of small RNA pathways.BMC Plant Biol. 2011 May 10;11:79. doi: 10.1186/1471-2229-11-79. BMC Plant Biol. 2011. PMID: 21569262 Free PMC article.
-
Root Development in Medicago truncatula: Lessons from Genetics to Functional Genomics.Methods Mol Biol. 2018;1822:205-239. doi: 10.1007/978-1-4939-8633-0_15. Methods Mol Biol. 2018. PMID: 30043307 Review.
Cited by
-
Comprehensive genomic analysis of the DUF4228 gene family in land plants and expression profiling of ATDUF4228 under abiotic stresses.BMC Genomics. 2020 Jan 3;21(1):12. doi: 10.1186/s12864-019-6389-3. BMC Genomics. 2020. PMID: 31900112 Free PMC article.
-
Heat Stress in Legume Seed Setting: Effects, Causes, and Future Prospects.Front Plant Sci. 2019 Jul 31;10:938. doi: 10.3389/fpls.2019.00938. eCollection 2019. Front Plant Sci. 2019. PMID: 31417579 Free PMC article.
-
A Comprehensive Analysis of Short Specific Tissue (SST) Proteins, a New Group of Proteins from PF10950 That May Give Rise to Cyclopeptide Alkaloids.Plants (Basel). 2025 Apr 3;14(7):1117. doi: 10.3390/plants14071117. Plants (Basel). 2025. PMID: 40219186 Free PMC article.
-
Specific tissue proteins 1 and 6 are involved in root biology during normal development and under symbiotic and pathogenic interactions in Medicago truncatula.Planta. 2021 Jan 2;253(1):7. doi: 10.1007/s00425-020-03538-4. Planta. 2021. PMID: 33387090
-
GCTTCA as a novel motif for regulating mesocarp-specific expression of the oil palm (Elaeis guineensis Jacq.) stearoyl-ACP desaturase gene.Plant Cell Rep. 2018 Aug;37(8):1127-1143. doi: 10.1007/s00299-018-2300-y. Epub 2018 May 22. Plant Cell Rep. 2018. PMID: 29789886
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

