Compound-specific isotope analysis resolves the dietary origin of docosahexaenoic acid in the mouse brain
- PMID: 28694298
- PMCID: PMC5625118
- DOI: 10.1194/jlr.D077990
Compound-specific isotope analysis resolves the dietary origin of docosahexaenoic acid in the mouse brain
Abstract
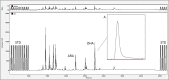
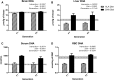
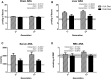
DHA (22:6n-3) may be derived from two dietary sources, preformed dietary DHA or through synthesis from α-linolenic acid (ALA; 18:3n-3). However, conventional methods cannot distinguish between DHA derived from either source without the use of costly labeled tracers. In the present study, we demonstrate the proof-of-concept that compound-specific isotope analysis (CSIA) by GC-isotope ratio mass spectrometry (IRMS) can differentiate between sources of brain DHA based on differences in natural 13C enrichment. Mice were fed diets containing either purified ALA or DHA as the sole n-3 PUFA. Extracted lipids were analyzed by CSIA for natural abundance 13C enrichment. Brain DHA from DHA-fed mice was significantly more enriched (-23.32‰ to -21.92‰) compared with mice on the ALA diet (-28.25‰ to -27.49‰). The measured 13C enrichment of brain DHA closely resembled the dietary n-3 PUFA source, -21.86‰ and -28.22‰ for DHA and ALA, respectively. The dietary effect on DHA 13C enrichment was similar in liver and blood fractions. Our results demonstrate the effectiveness of CSIA, at natural 13C enrichment, to differentiate between the incorporation of preformed or synthesized DHA into the brain and other tissues without the need for tracers.
Keywords: fatty acid; omega-3 fatty acids; stable isotope analysis.
Copyright © 2017 by the American Society for Biochemistry and Molecular Biology, Inc.
Conflict of interest statement
R.P.B. holds a Canada Research Chair in Brain Lipid Metabolism and has received research grants from Bunge Ltd., Arctic Nutrition, the Dairy Farmers of Canada, and Nestle Inc., as well as travel support from Mead Johnson and mass spectrometry equipment and support from Sciex. R.P.B. is on the executive committee of the International Society for the Study of Fatty Acids and Lipids and held a meeting on behalf of Fatty Acids and Cell Signaling, both of which rely on corporate sponsorship. R.P.B. has given expert testimony in relation to supplements and the brain. R.P.B. also provides complementary FA analysis for farmers, food producers, and others involved in the food industry, some of whom provide free food samples. V.G. holds a Canadian Institutes of Health Research Canada Graduate Scholarship.
Figures




References
-
- Martinez M. 1992. Abnormal profiles of polyunsaturated fatty acids in the brain, liver, kidney and retina of patients with peroxisomal disorders. Brain Res. 583: 171–182. - PubMed
-
- Bazinet R. P., and Layé S.. 2014. Polyunsaturated fatty acids and their metabolites in brain function and disease. Nat. Rev. Neurosci. 15: 771–785. - PubMed
-
- Kim H. Y. 2007. Novel metabolism of docosahexaenoic acid in neural cells. J. Biol. Chem. 282: 18661–18665. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

