Rapid outbreak sequencing of Ebola virus in Sierra Leone identifies transmission chains linked to sporadic cases
- PMID: 28694998
- PMCID: PMC5499387
- DOI: 10.1093/ve/vew016
Rapid outbreak sequencing of Ebola virus in Sierra Leone identifies transmission chains linked to sporadic cases
Abstract
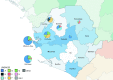
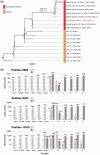
To end the largest known outbreak of Ebola virus disease (EVD) in West Africa and to prevent new transmissions, rapid epidemiological tracing of cases and contacts was required. The ability to quickly identify unknown sources and chains of transmission is key to ending the EVD epidemic and of even greater importance in the context of recent reports of Ebola virus (EBOV) persistence in survivors. Phylogenetic analysis of complete EBOV genomes can provide important information on the source of any new infection. A local deep sequencing facility was established at the Mateneh Ebola Treatment Centre in central Sierra Leone. The facility included all wetlab and computational resources to rapidly process EBOV diagnostic samples into full genome sequences. We produced 554 EBOV genomes from EVD cases across Sierra Leone. These genomes provided a detailed description of EBOV evolution and facilitated phylogenetic tracking of new EVD cases. Importantly, we show that linked genomic and epidemiological data can not only support contact tracing but also identify unconventional transmission chains involving body fluids, including semen. Rapid EBOV genome sequencing, when linked to epidemiological information and a comprehensive database of virus sequences across the outbreak, provided a powerful tool for public health epidemic control efforts.
Keywords: Ebola virus; evolution; outbreak sequencing; transmission.
Figures



References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

