Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial mesenchymal transition by sponging miR-150-5p
- PMID: 28697764
- PMCID: PMC5504775
- DOI: 10.1186/s12943-017-0685-9
Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial mesenchymal transition by sponging miR-150-5p
Erratum in
-
Erratum to: Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial mesenchymal transition by sponging miR-150-5p.Mol Cancer. 2017 Aug 29;16(1):144. doi: 10.1186/s12943-017-0716-6. Mol Cancer. 2017. PMID: 28851432 Free PMC article. No abstract available.
Abstract
Background: The function of a new long non-coding RNA linc00673 remains unclear. While identified as an oncogenic player in non-small cell lung cancer (NSCLC), linc00673 was found to be anti-oncogenic in pancreatic ductal adenocarcinoma (PDAC). However whether linc00673 regulated malignancy and epithelial mesenchymal transition (EMT) has not been characterized.
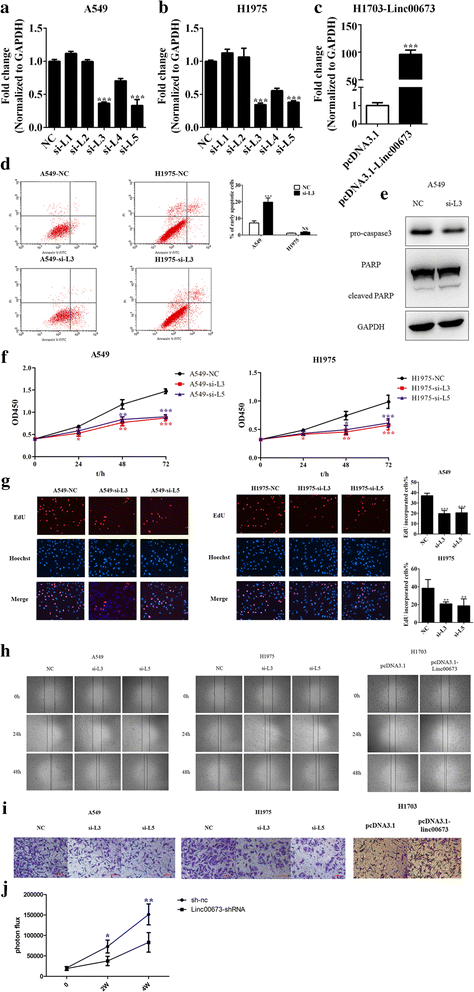
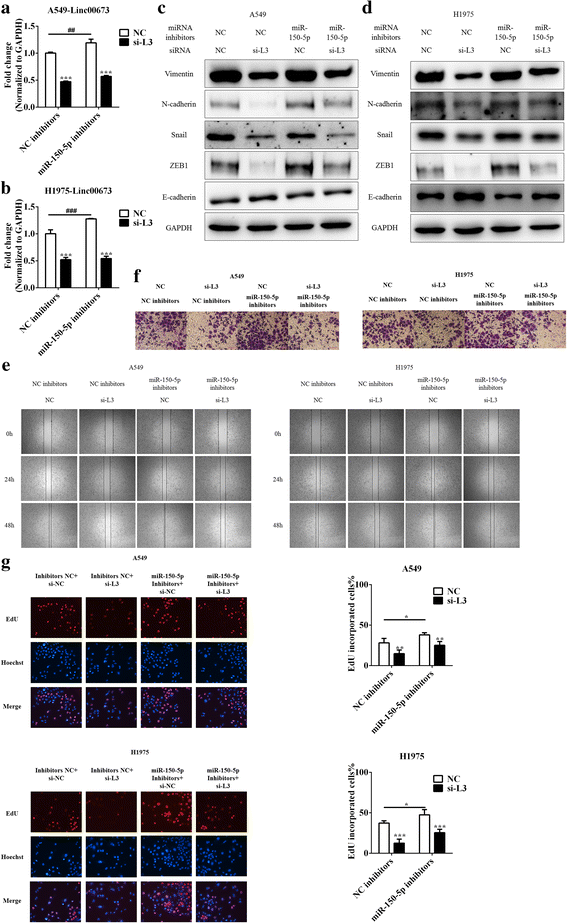
Methods: Cell proliferation was assessed using CCK-8 and EdU assays, and cell migration and invasion were assessed using scratch assays and transwell invasion assays. Epithelial mesenchymal transition was examined using western blot, qRT-PCR and immunofluorescence staining. Interaction between miRNA and linc00673 was determined using luciferase reporter assays. In vivo experiments were performed to assess tumor formation. In addition, the expression data of NSCLC specimens of TCGA and patient survival data were utilized to explore the prognostic significance of linc00673.
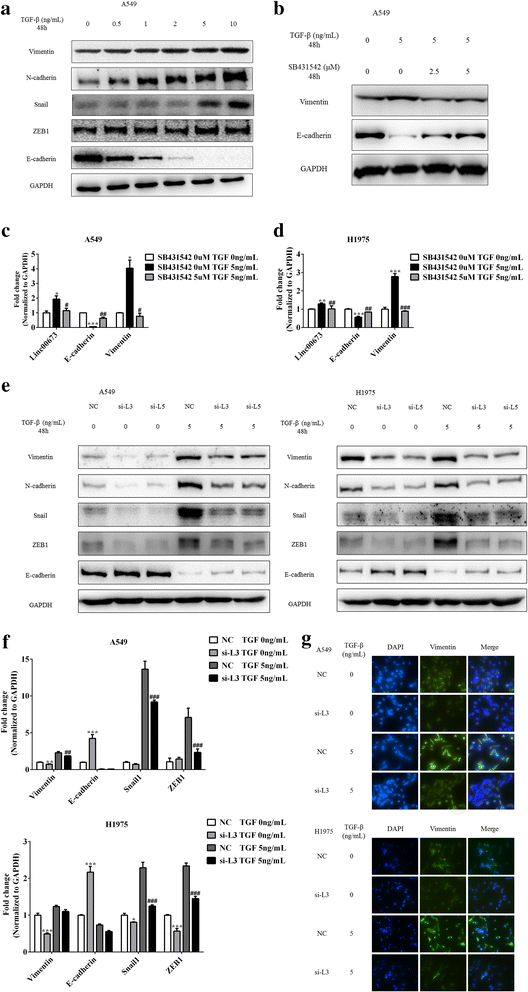
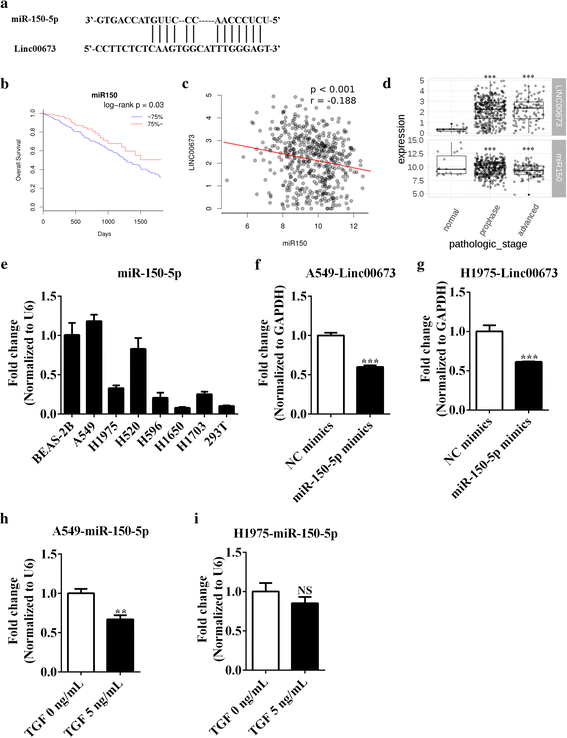
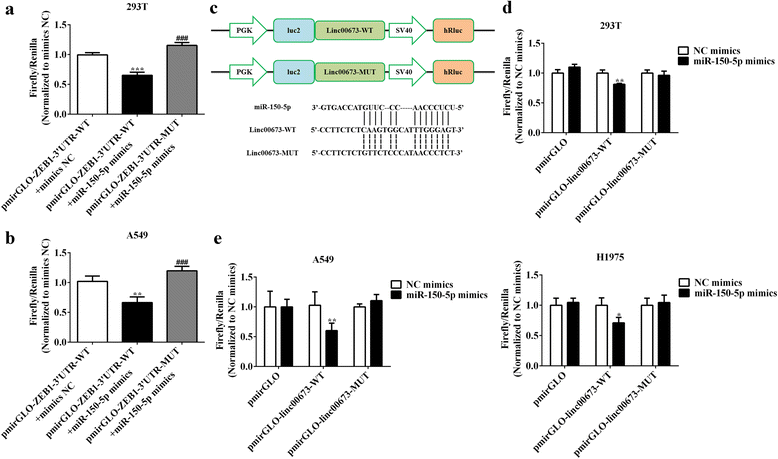
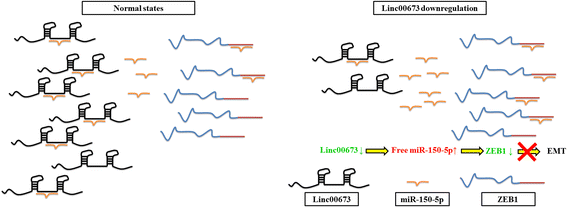
Results: In the present study, we found high linc00673 expression was associated with poor prognosis of NSCLC patients. In vitro experiments showed linc00673 knockdown reversed TGF-β induced EMT, and miR-150-5p was predicted to target linc00673 through bioinformatics tools. Overexpression of miR-150-5p suppressed lin00673's expression while inhibition of miR-150-5p led to significant upregulation of lin00673, suggesting that linc00673 could be negatively regulated by miR-150-5p, which was further confirmed by the inverse correlation between linc00673 and miR-150-5p in NSCLC patients' specimen. Furthermore, we proved that miR-150-5p could directly target linc00673 through luciferase assay, so linc00673 could sponge miR-150-5p and modulate the expression of a key EMT regulator ZEB1 indirectly. In addition, miR-150-5p inhibition abrogated linc00673 silence mediated proliferation, migration, invasion and EMT suppressing effect. Moreover, the inhibition of linc00673 significantly attenuated the tumorigenesis ability of A549 cells in vivo.
Conclusions: We validated linc00673 as a novel oncogenic lncRNA and demonstrated the molecular mechanism by which it promotes NSCLC, which will advance our understanding of its clinical significance.
Keywords: Competing endogenous RNA; Epithelial mesenchymal transition; Non-small cell lung cancer; linc00673; miR-150-5p.
Conflict of interest statement
Ethics approval and consent to participate
All animal experiments were approved by the ethical review committee from Zhejiang University School of Medicine.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures

References
-
- Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447:799–816. doi: 10.1038/nature05874. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

