Crystal structure of CO-bound cytochrome c oxidase determined by serial femtosecond X-ray crystallography at room temperature
- PMID: 28698372
- PMCID: PMC5544322
- DOI: 10.1073/pnas.1705628114
Crystal structure of CO-bound cytochrome c oxidase determined by serial femtosecond X-ray crystallography at room temperature
Abstract
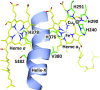
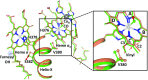
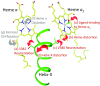
Cytochrome c oxidase (CcO), the terminal enzyme in the electron transfer chain, translocates protons across the inner mitochondrial membrane by harnessing the free energy generated by the reduction of oxygen to water. Several redox-coupled proton translocation mechanisms have been proposed, but they lack confirmation, in part from the absence of reliable structural information due to radiation damage artifacts caused by the intense synchrotron radiation. Here we report the room temperature, neutral pH (6.8), damage-free structure of bovine CcO (bCcO) in the carbon monoxide (CO)-bound state at a resolution of 2.3 Å, obtained by serial femtosecond X-ray crystallography (SFX) with an X-ray free electron laser. As a comparison, an equivalent structure was obtained at a resolution of 1.95 Å, from data collected at a synchrotron light source. In the SFX structure, the CO is coordinated to the heme a3 iron atom, with a bent Fe-C-O angle of ∼142°. In contrast, in the synchrotron structure, the Fe-CO bond is cleaved; CO relocates to a new site near CuB, which, in turn, moves closer to the heme a3 iron by ∼0.38 Å. Structural comparison reveals that ligand binding to the heme a3 iron in the SFX structure is associated with an allosteric structural transition, involving partial unwinding of the helix-X between heme a and a3, thereby establishing a communication linkage between the two heme groups, setting the stage for proton translocation during the ensuing redox chemistry.
Keywords: X-ray free electron laser; bioenergetics; crystallography; cytochrome c oxidase; serial femtosecond crystallography.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Yoshikawa S, Muramoto K, Shinzawa-Itoh K. Proton-pumping mechanism of cytochrome C oxidase. Annu Rev Biophys. 2011;40:205–223. - PubMed
-
- Yoshikawa S, Shimada A. Reaction mechanism of cytochrome c oxidase. Chem Rev. 2015;115:1936–1989. - PubMed
-
- Kaila VR, Verkhovsky MI, Wikström M. Proton-coupled electron transfer in cytochrome oxidase. Chem Rev. 2010;110:7062–7081. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

