Repair or destruction-an intimate liaison between ubiquitin ligases and molecular chaperones in proteostasis
- PMID: 28699655
- PMCID: PMC5601288
- DOI: 10.1002/1873-3468.12750
Repair or destruction-an intimate liaison between ubiquitin ligases and molecular chaperones in proteostasis
Abstract
Cellular differentiation, developmental processes, and environmental factors challenge the integrity of the proteome in every eukaryotic cell. The maintenance of protein homeostasis, or proteostasis, involves folding and degradation of damaged proteins, and is essential for cellular function, organismal growth, and viability . Misfolded proteins that cannot be refolded by chaperone machineries are degraded by specialized proteolytic systems. A major degradation pathway regulating cellular proteostasis is the ubiquitin (Ub)/proteasome system (UPS), which regulates turnover of damaged proteins that accumulate upon stress and during aging. Despite a large number of structurally unrelated substrates, Ub conjugation is remarkably selective. Substrate selectivity is mainly provided by the group of E3 enzymes. Several observations indicate that numerous E3 Ub ligases intimately collaborate with molecular chaperones to maintain the cellular proteome. In this review, we provide an overview of specialized quality control E3 ligases playing a critical role in the degradation of damaged proteins. The process of substrate recognition and turnover, the type of chaperones they team up with, and the potential pathogeneses associated with their malfunction will be further discussed.
Keywords: CHIP; E3 ligase; aging; chaperone; proteostasis; quality control; ubiquitin.
© 2017 The Authors. FEBS Letters published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Figures

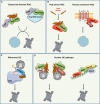
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

